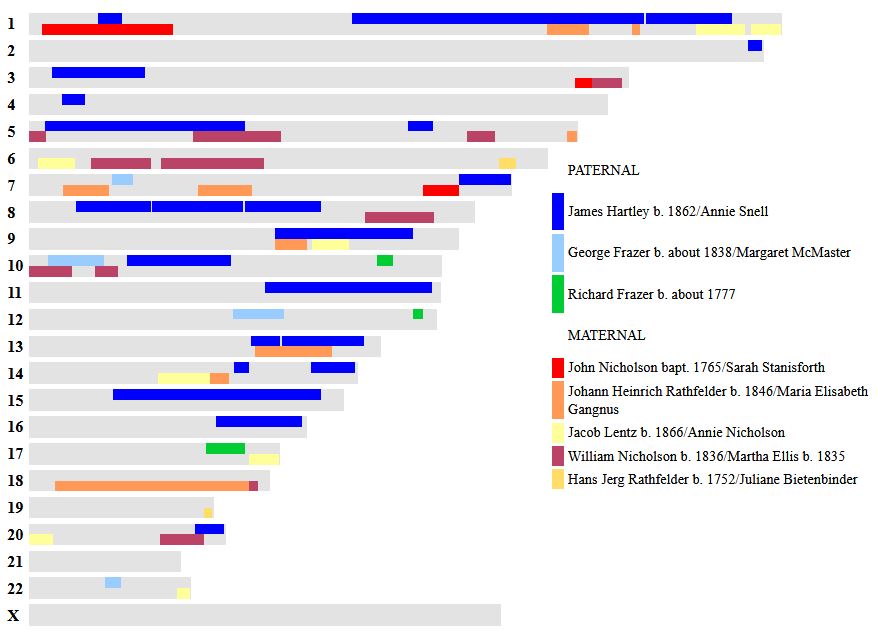
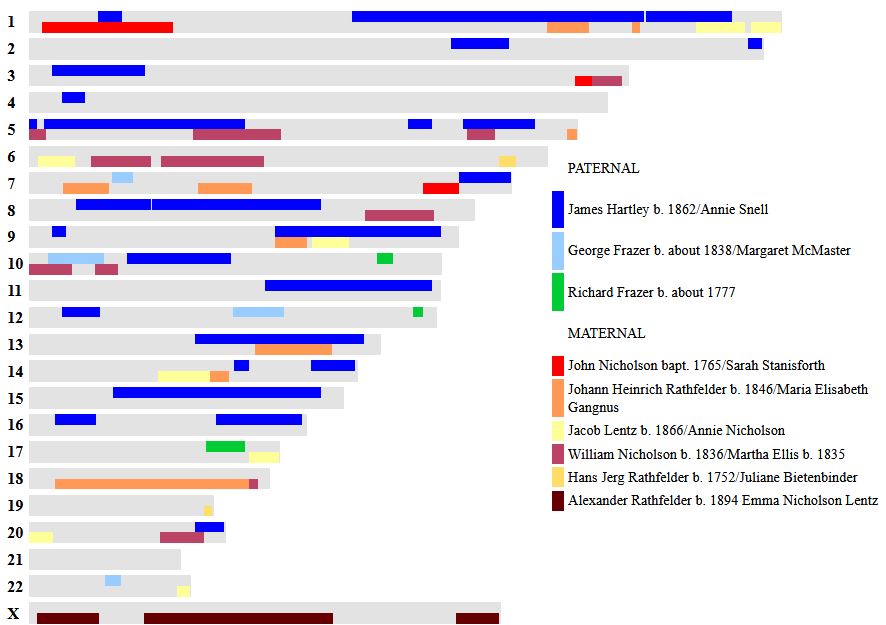
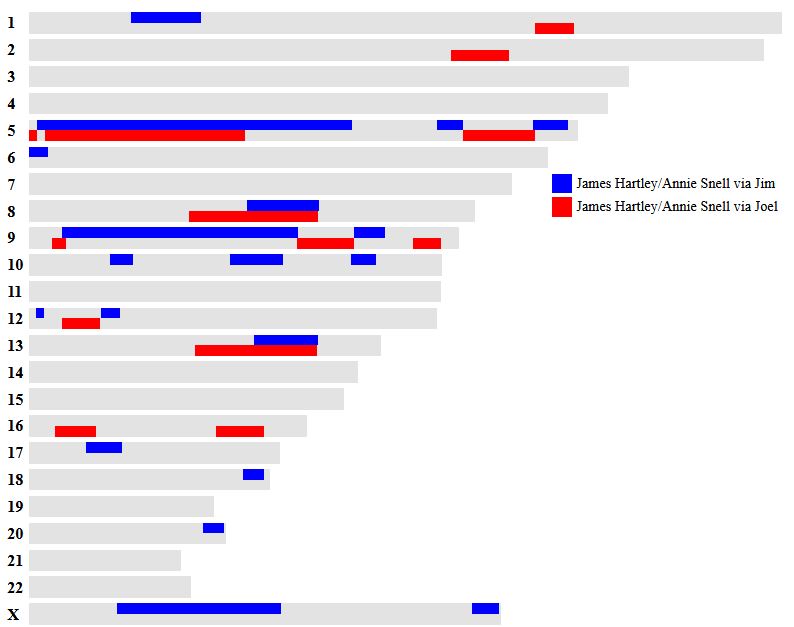
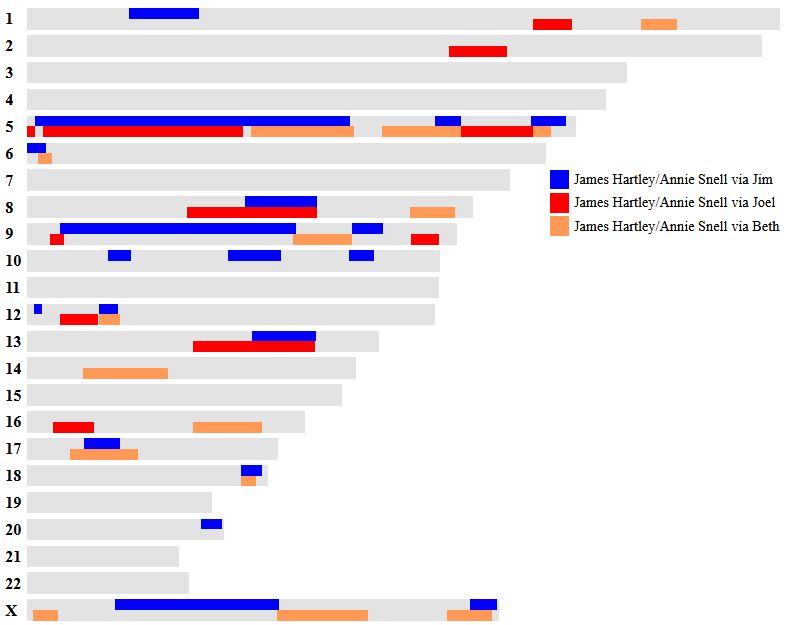
In my previous Blog, I looked at Whit Athey’s Principle 3 for my mom, my 4 siblings and myself. Based on that Priniciple and the previous 2, I phased our DNA up to a point. The next step in the phasing has to do with patterns.
Patterns
The patterns I am talking about are the patterns that the five siblings receive from either their mother or father. For example, an AAAAB pattern means that the first 4 siblings received the same allele and the fifth sibling received a different one. I had mentioned previously that the patterns should be in this form:
- AAAAA
- AAAAB
- AAABA
- AAABB
- AABAA
- AABAB
- AABBB
- ABAAA
- ABAAB
- ABABB
- ABBBB
The first situation is a special case as this situation can happen within the other patterns ‘by accident’ as Athey puts it.
AAAAB Dad pattern
First I’ll look at a query to find an AAAAB pattern.
That Query results in this:
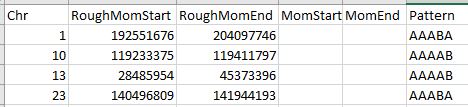
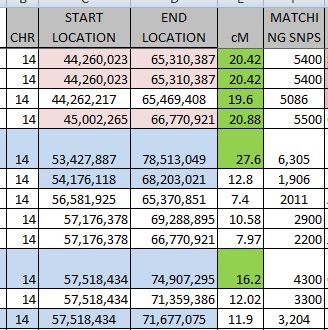
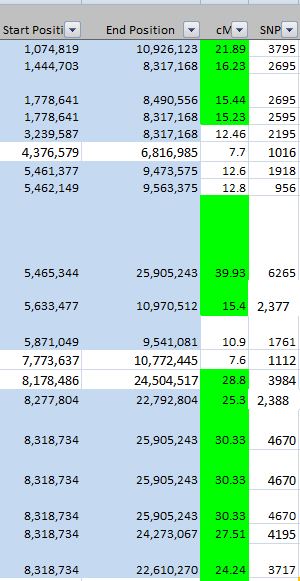
Except there are actually over 8,000 lines. I summarized the rough starts and stops in an Excel Spreadsheet:
This part can get a bit tedious. In Chromosome 2, I noted a possible break between 89 and 96M, so I’ll need to keep an eye on that. Highlighted in yellow are single patterns which may or may not be significant.
quality check
I took my AAAAB Query results and put them into an Excel spreadsheet. Then I subtracted the previous position number from the current position number to see where there were gaps. Then I filtered the gap to 1,000,000 or more positions:
This is my gap analysis. I highlighted the 7 million position gap where I put in an extra segment on the AAAAB pattern. This points out some of the single AAAAB patterns also.
mapping the initial results
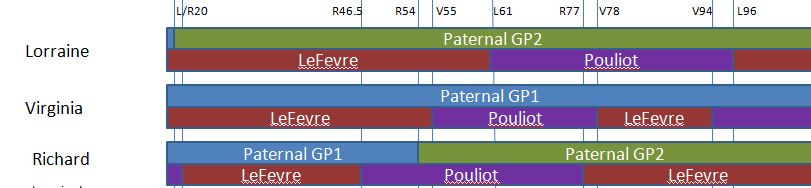
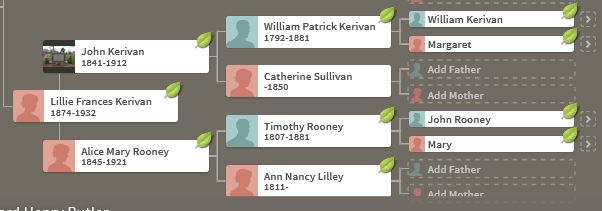
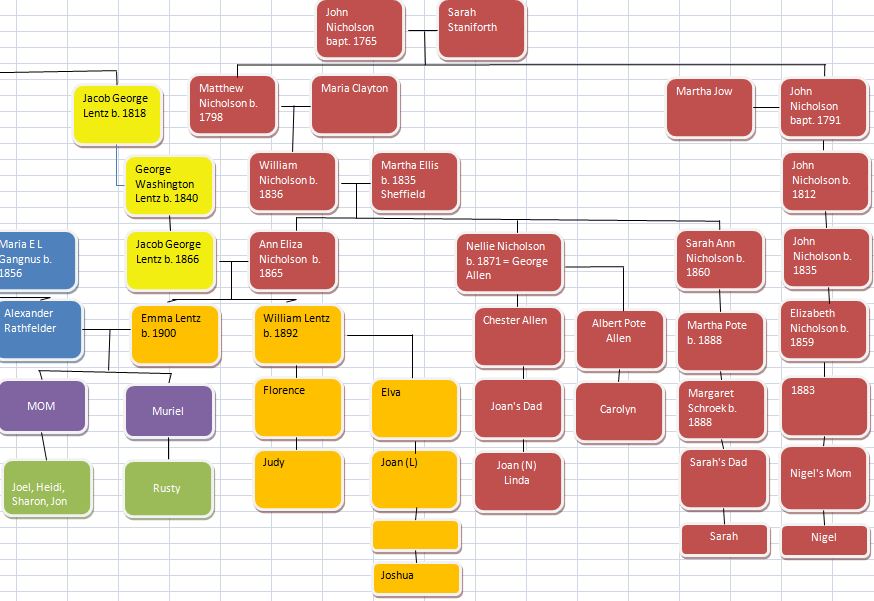
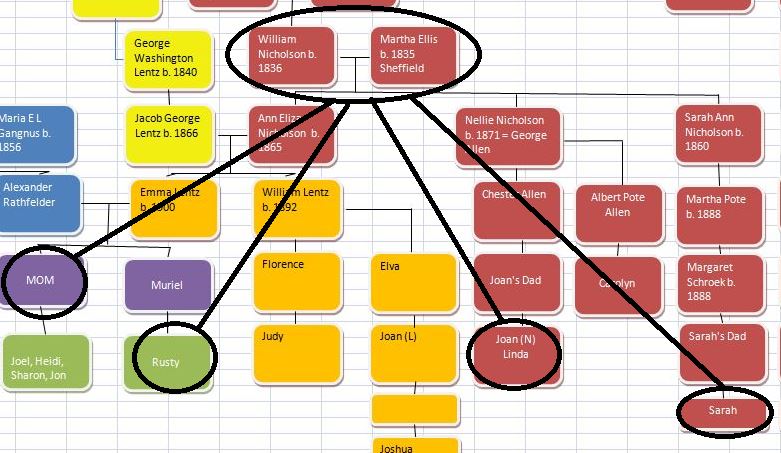
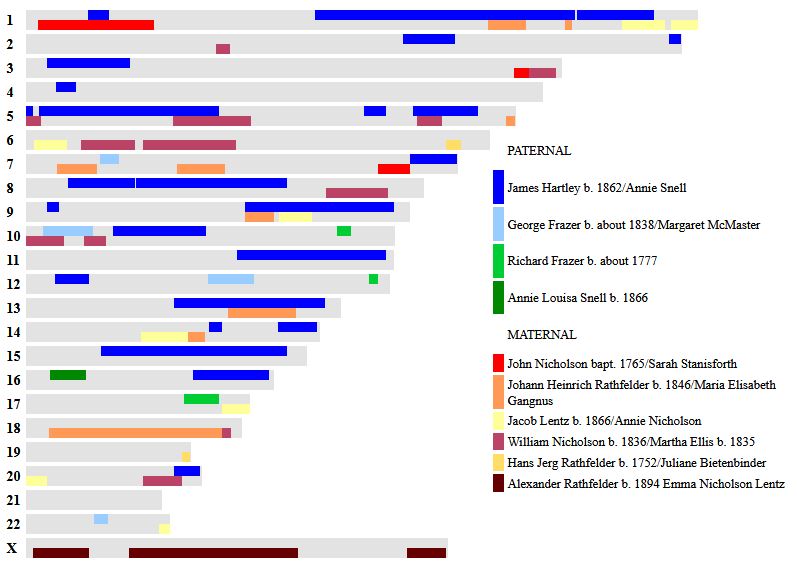
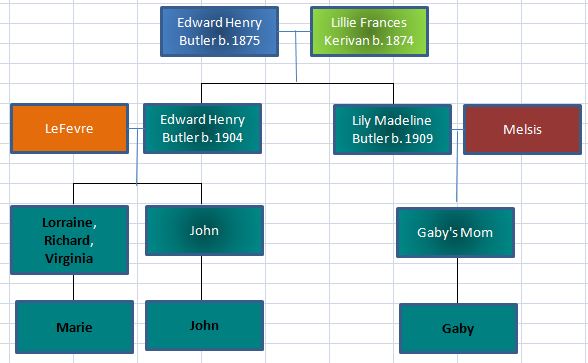
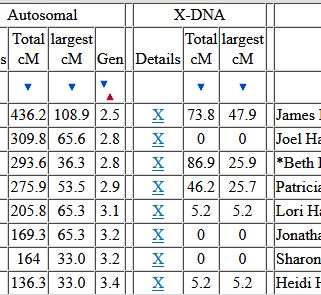
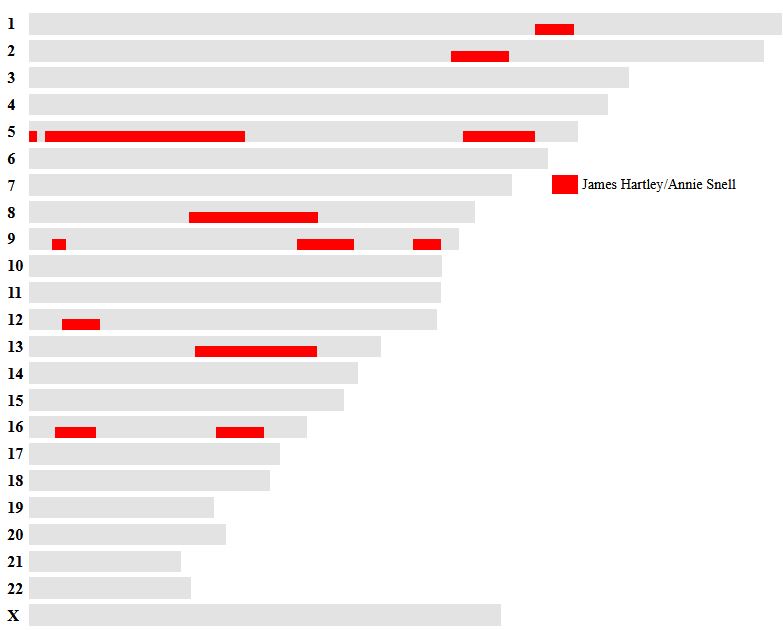
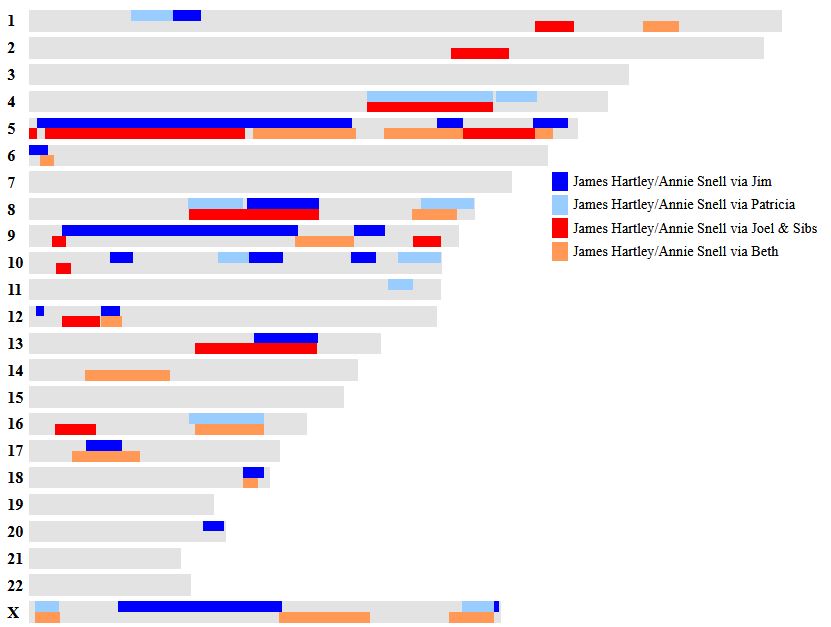
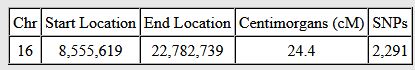
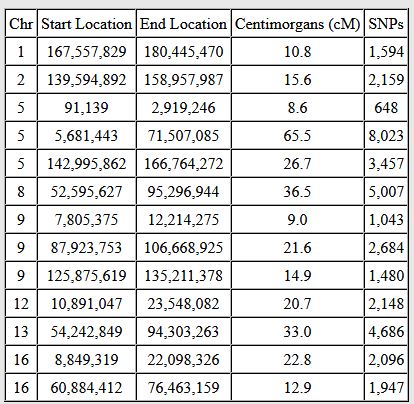
Let’s look at Chromosome 13 between 28 and 87M. With an AAAAB pattern, that means that Joel, Sharon, Heidi and Jon match the same paternal grandparent. Lori matches a different one. However, we don’t know which paternal grandparent without a reference cousin. Fortunately, I have one. He is my dad’s first cousin. He would match on my paternal grandfather’s side. That grandfather is James Hartley, b. 1891:
Paternal cousin Jim matches the 5 siblings here:
As you may guess, Lori is on the bottom (#5). She has a crossover at about 85.5M according to Gedmatch. That means that before 85.5M she is matching on my father’s mother’s side: Marion Frazer. So, if I wanted to, I could start to map Chromosome 13. From 28 to 87M, I could say that 4 siblings got their DNA from their paternal Hartley grandfather and one sibling, Lori got hers from her paternal Frazer grandmother.
Further, I would expect an AAAAA pattern starting at 87M based on the gedmatch browser results above. The bad thing about an AAAAA pattern is that there is some missing DNA for the other grandparent. In this case, the Frazer DNA is lost on the right side of the map below. Another point is that these patterns change one letter at a time. So it makes sense that an AAAAB would go to an AAAAA. For example, an AAAAA would never go directly to an ABABA.
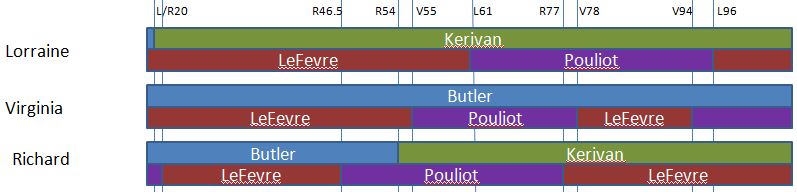
Here is a paternal only map of Chromosome 13 based on our very initial results:
aaaab Mom pattern
I notice that the formula that I used to find the AAAAB Dad pattern, I can move over to the mom side. So I might as well do this while I’m thinking of AAAAB pattenrs and put the results in Excel.
I randomly used Heidi as the ‘A’. So Lori not matching Heidi becomes the ‘B’. The results for this maternal query was much smaller with only 189 lines.
This was a lot easier. The Mom and Dad Patterns don’t interrelate with each other, so I have them on separate worksheets. Note that there is the same AAAAB pattern in the same starting place on Chromosome 13 as there was on the paternal side. This is a coincidence and the starting spot is a coincidence. This is just a rough number now and may be refined later. I could make a map of this also.
Here is a cousin on my mom’s father’s side:
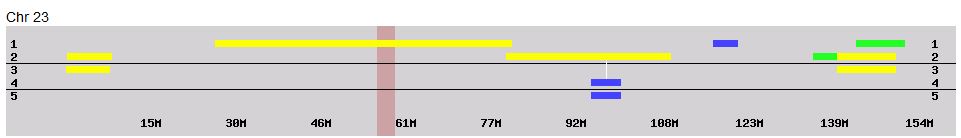
Here she matches Joel, Sharon, Heidi and Lori from about 74-99M. Here is a map drawn on the Gedmatch browser and raw data phasing:
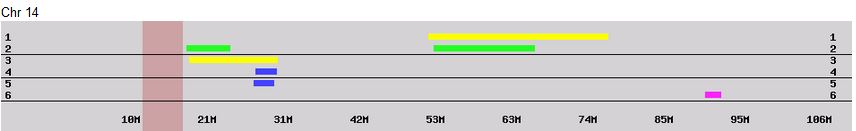
This shows what a AAAAB pattern looks like that is both paternal and maternal between 28 and 45M. I also show two crossovers for Lori: (Frazer to Hartley) and (Rathfelder to Lentz). In addition, Jon has to have a crossover from Rathfelder to Lentz and Lori has to have another crossover from Lentz to Rathfelder somewhere in the white spaces. There is a reason that I could tell the maternal A’s of the AAAAB pattern were Rathfelder even though our cousin match did not overlap that area. It is because the patterns do not change that fast as I explained above.
Now that I know which sibling has one of the paternal crossovers I can mark it on the Dad Pattern Spreadsheet:
I name the crossover column in the spreadsheet for the end of the pattern position, so it will be clear where it is. This is the ultimate goal of the process: to find the crossover locations and assign them to the siblings. Once this is done a map may be drawn for all the siblings.
The Next Step
In the next step, I could fill in the missing alleles between the Start and End positions of the AAAAB patterns. Here is how that will be done:
The highlighted row is where the AAAAB Pattern starts. Basically, what will happen is if there is at least one Allele in the first four positions, I will be able to fill in any of the other alleles in those first four positions with the same allele. However, in the last row, for example, there is just one G in the last position. We don’t know if the other four alleles will be a G or another letter. The row that has TTT??. We know that we can fill in the fourth T to TTTT?. However, the last allele we don’t know if it will be a T again or a different alelle. So we also need to leave that space blank.
However, I want to make sure I have all my patterns right, so I will look at all the patterns first and reconcile them.
AAABA Pattern
If I drew my map correctly above, I will be expecting Lori to have a maternal AAABA pattern on Chromosome 13. This should change to an AAABB pattern at about position 95M. I’m already on a maternal query, so I’ll start there.
I used Heidi again as the A. Now Jon is the B that is different than Heidi. I was surprised with the results as I only had this maternal pattern in Chromosome 1 and 23:
My prediction of a Chromosome 13 AAABA Pattern did not come true. I wonder what went wrong?
Paternal AAABA Pattern
Here is a partial summary of the Paternal AAABA Pattern:
On Chromosome 11, we see the AAABA pattern twice with an AAAAB pattern in-between. To go from an AAAAB to AAABA there has to be a transition pattern: either AAAAA or AAABB. Hopefully this prediction will be correct! That leads me to the AAABB pattern.
AAABB
This pattern requires a slight modification of my previous query:
This pattern is adjacent to the AAABA Pattern, so I will be able to assign some crossovers:
These crossovers belong to Jon and Lori as Jon is in the next to the last position of the patterns and Lori is in the last position of the patterns. Note that in Chromosome 19, Lori goes from an AAABA to an AAABB at about 5M. However, there is a rogue AAABB in the AAABA pattern at around 3M. That could be due to a misread or a mutation. I’m not sure. Jon has a crossover on Chromosome 8. These are all Lori and Jon crossovers, due to the positions of the pattern changes we are looking at. The changes are all in the last two positions.
AAABB Maternal
I’m still getting very few crossovers here. I’m not sure why:
I’m not sure why the maternal side is not keeping up with the paternal. I have no crossovers here yet.
AABAA
Following my alphabetical reasoning, AABAA is my next pattern. I’ll start with the maternal:
I changed to having my [Joel’s] allele the ‘A’ in the Pattern. The results look right:
It seemed like there was a break in Chromosome 5 between 46 and 50M.
AABAA Paternal
On Chromosome 5, there was a gap similar to the one on the Maternal side.
Centromeres
According to ISOGG, these are the Build 37 Centromeres:
This is good information to have. I assume that the Centromere is not counted, so I will ignore the Chromosome 5 missing area and make a note that the centromere is there. This also makes a difference on all the results.
AABAB: Are We There Yet?
Here are Heidi’s first crossovers. I’ve also heard of crossovers referred to as cut points. I am noting where the centromere is – though not quite spelling it correctly above.
Here is the Maternal AABAB. I am still annoyed that there are so few patterns. They seem to be missing for some reason:
I suppose, if this trend continues, I could do the project over and add in my mother’s and my FTDNA raw DNA results.
AABBA
I didn’t find any AABBA Patterns on the maternal side. However, that was with a query using my results as A. However, from my previous Blog, I recall this chart:
This showed that on the Mom side, Jon and Lori had the most alleles. I’ll run the query again this way:
Still no patterns.
Here are some more Dad Patterns:
However, there are a few problems. Chromosome 17 is missing a pattern. I can solve this by looking at the original table.
Here the pattern is AABAA.
The next problem is that there are two patterns in one spot on Chromosome 22. I ran pattern AAABA again and see it should have ended earlier:
Here is the right answer below that also shows a Heidi crossover at that location:
AABBB
Paternal
Maternal side
Still nothing.
ABAAA
Maternal side
Paternal
This side had more patterns.
ABAAB
Had several of ABAAB Patterns on the dad side, but only one on the mom side. I think that there is a fill-in step that fills in the mom side from the dad side that may correct this later.
ABABA, ABABB, ABBAA
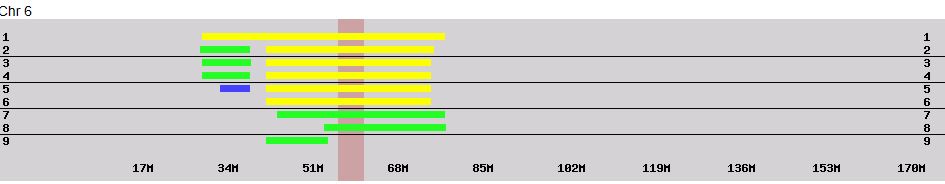
I did notice a Dad Pattern discrepancy on Chromosome 6:
There are three single patterns, I figured were discrepancies. However, there appeared to be a longer AAABB Pattern within the ABBAA Pattern. This is where it helps to look at the raw data.
The blue section is the start of the AAABB rogue pattern that I had. However, a closer examination reveals that this pattern is not continuous from position 30514810 to 30594827. Between those two there are a lot of ABBAA patterns. This is clear at position 30544401. However, this is also clear wherever the first 2 alleles are different. For example, on the last line, I see GT???. This will be filled in with GTTAA as this is within the ABBAA Pattern area. So what happened was that there were two single AAABB patterns. When I did the query for these, it looked like the pattern was continuous, but it was not. Based on the above, I’ve modified my Dad Pattern Spreadsheet to show two single discrepancies:
I won’t overwrite this information, but I will keep it in mind for later in case it is important. If this was a real crossover, it would be mine. However, crossovers in the middle of a chromosome don’t change that fast for one person on one copy of their chromosome.
Some of the Dad Pattern Crossovers are starting to fill in:
Starts and Ends of Chromosomes
At some point, it is important to know where the Chromosomes start and end. The testing companies don’t always start at the beginning positions of each chromosome. The ends are different also based on the lengths of the chromosomes.
I was able to find what I was looking for using a min/max Query in Access. I took my table with the 900,000 plus alleles and made a query that looks like this:
When I run the Query, I get this helpful table:
This tells me the start and end locations for each of the chromosomes that I am looking at.
I put this into Excel and highlighted the information in purple. Then I sorted it into my mom and dad pattern spreadsheets:
Now, I can tell that I am near the beginning and the end of Chromosome 20 with the pattern locations. However, on Chromosomes 21 and 22, there is still room for more at the beginning of those Chromosomes. As the Chromosome 20 patterns are complete, this also tells me that my sister Lori has no paternal crossover on Chromosome 20.
ABBAB, ABBBA, and ABBBB
These are the last three patterns, not counting AAAAA. I finally have one crossover on the maternal side. It is on the X Chromosome:
I have a mess to clean up on Paternal Chromosome 2 :
There appear to be two patterns occupying the same space between 123 and 128M, which is not good. I’ll take a look at my Table: It appears that the AAAAB at 127,841,390 is a one-time occurrence. Here is my correction:
Note that there is still a gap at AAAAB. There may be an AAAAA Pattern stuck in there.
Lessons Learned and Next Up
- It is good to document the process in case something goes wrong
- The start and end points are needed for each chromosome
- The start and end for each centromere is needed also
- Attention is needed for the location of each crossover and who it goes to as this is a main point of all the work.
- Changes along each copy of the chromosome are gradual. They happen one at a time and those one at a time changes correspond to siblings.
- Next up is filling in the blanks. That was discussed briefly in this Blog.