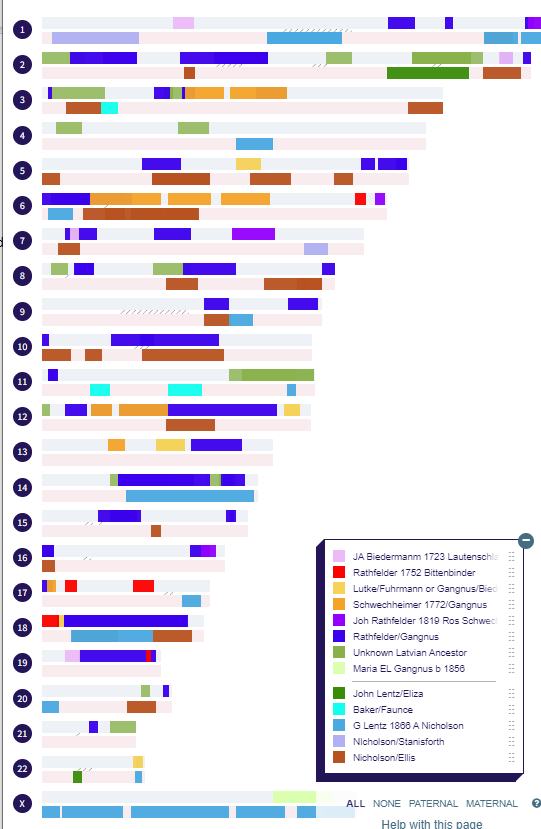
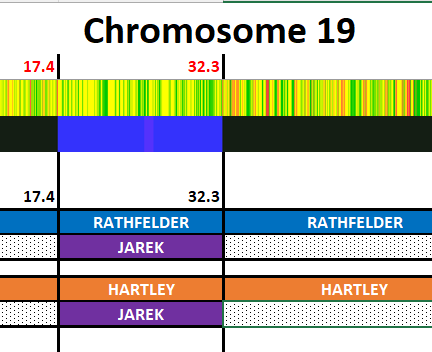
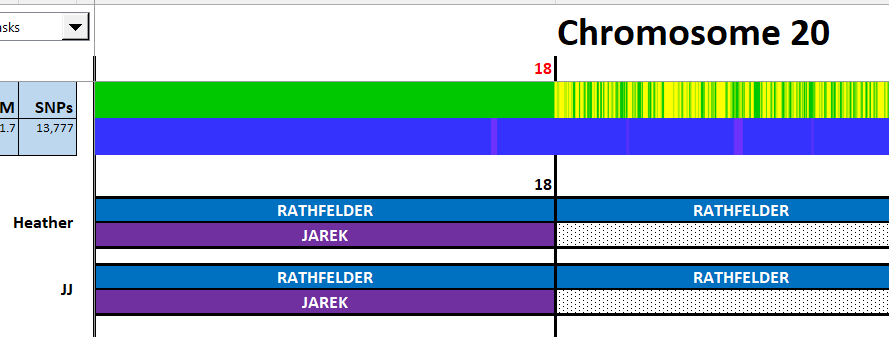
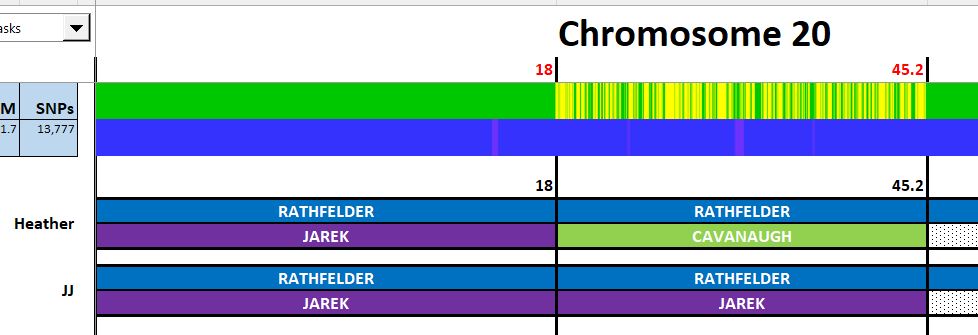
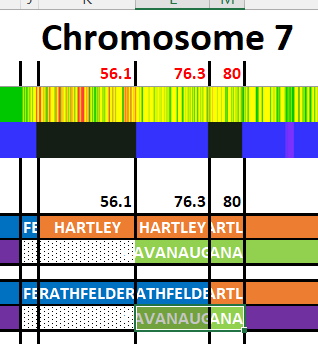
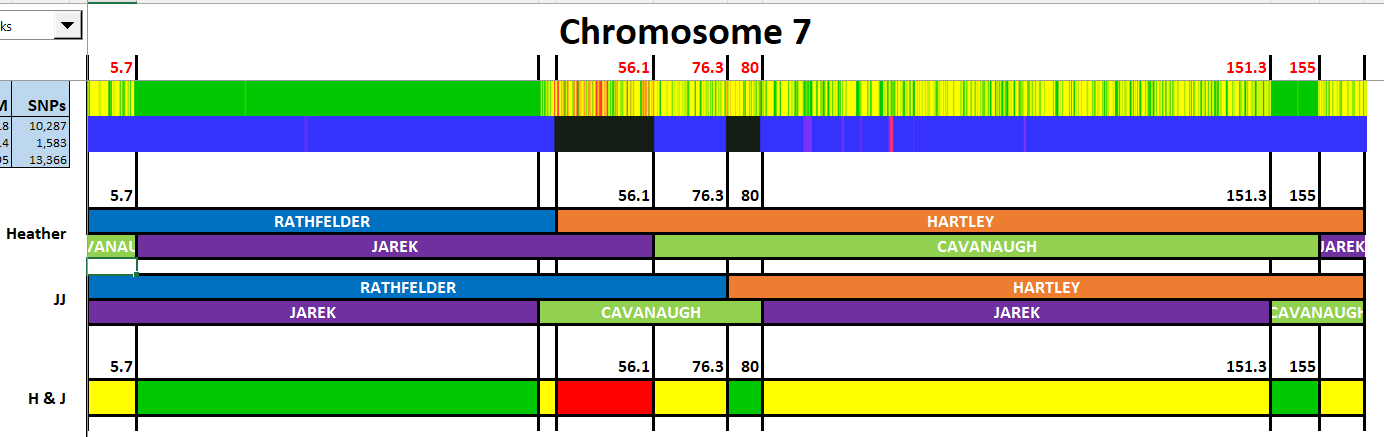
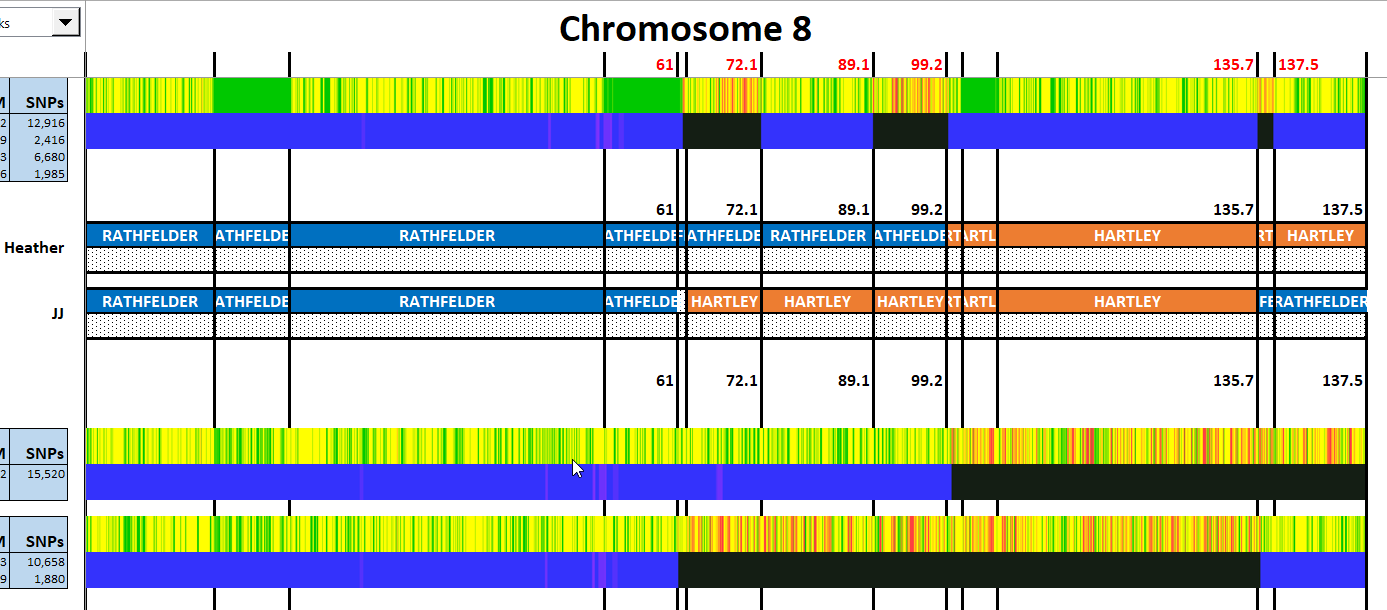
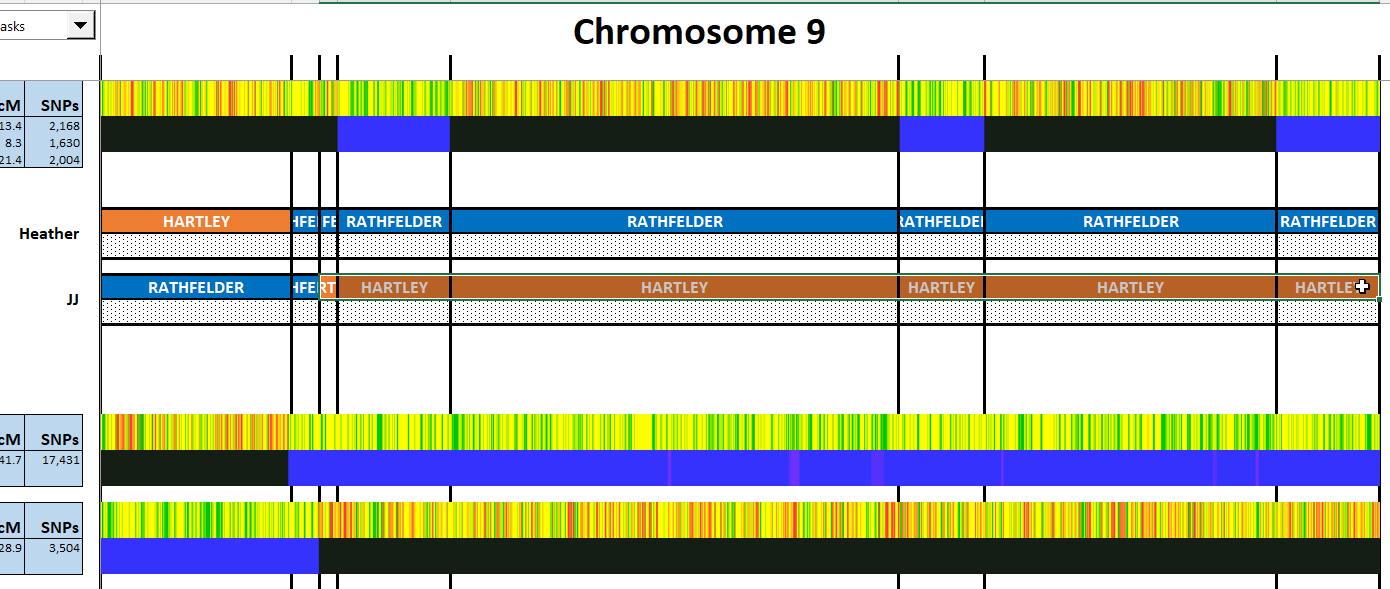
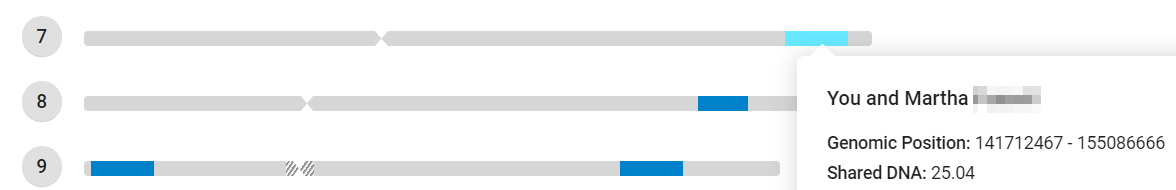I have made a few isolated attempts at Visual Phasing for my two children. However, I have not used the Fox Spreadsheet up to this point. This spreadsheet is not supposed to work well with the online version of Excel, but I am trying it nonetheless.
Getting Started
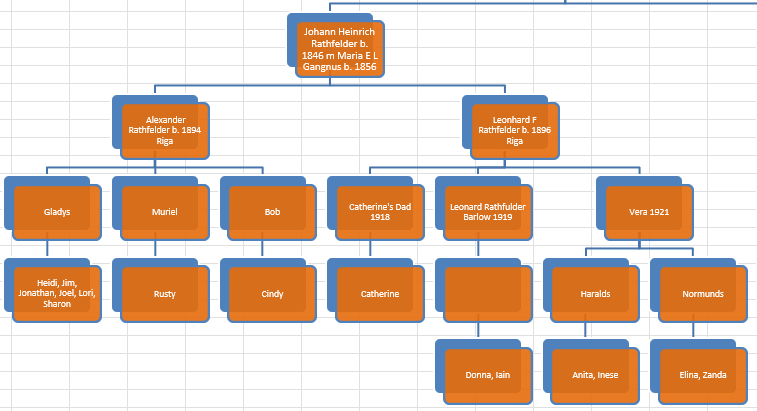
I downloaded my two children’s information from Gedmatch in addition to three “cousins” matches. One cousin is my mother. She is the secret weapon as her matches should automatically phase my children’s patermal side. The other two matches were two second cousins once removed. One was on the Hartley side and one was on their Jarek side. I don’t think that I have cousin matches on the Cavanaugh side. If I do, I don’t think that there is a good match at Gedmatch.
I looked at Steven Fox’s intstructions. In order to get a download from Gedmatch, it required my pushing a ‘compare’ button from time to time to get around the Captcha feature at Gedmatch.
Chromosome 1
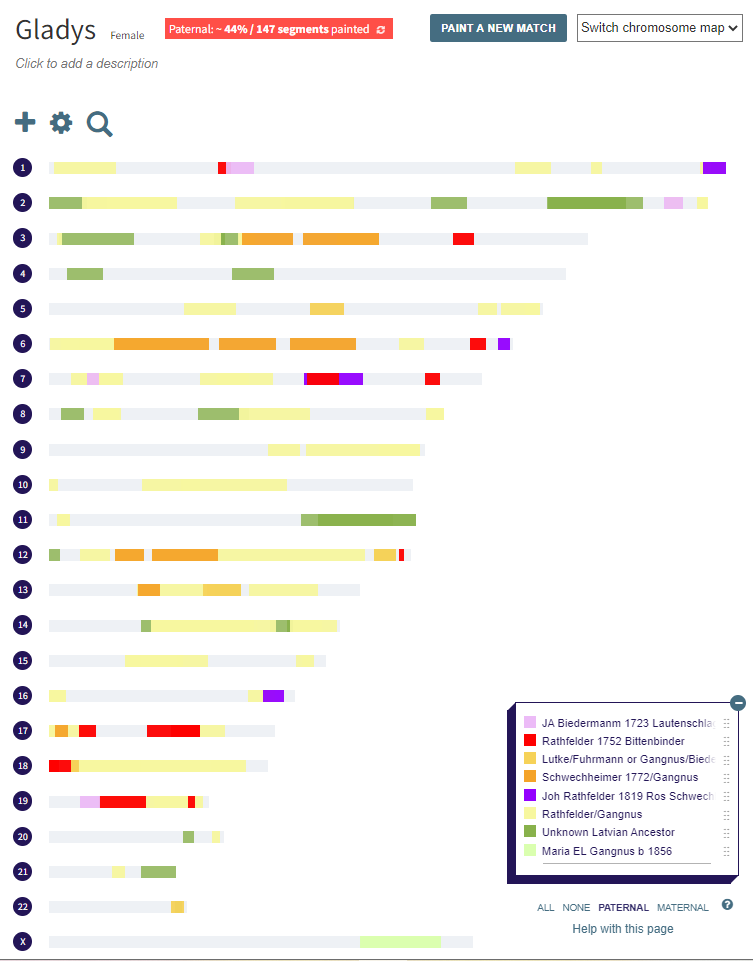
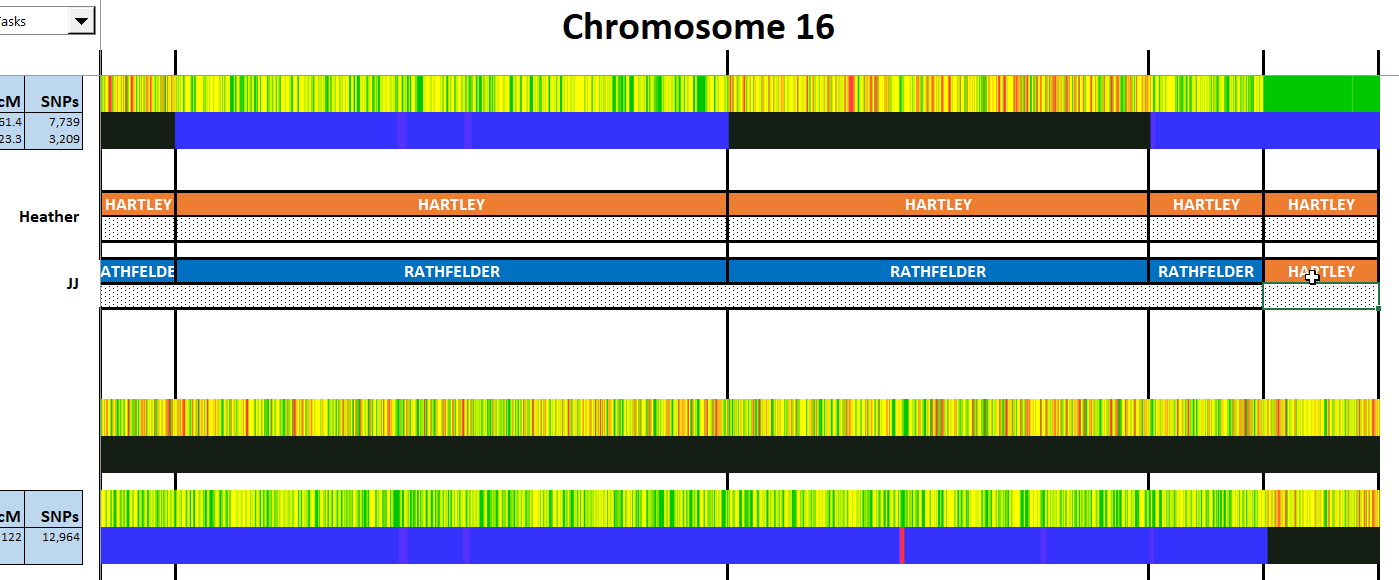
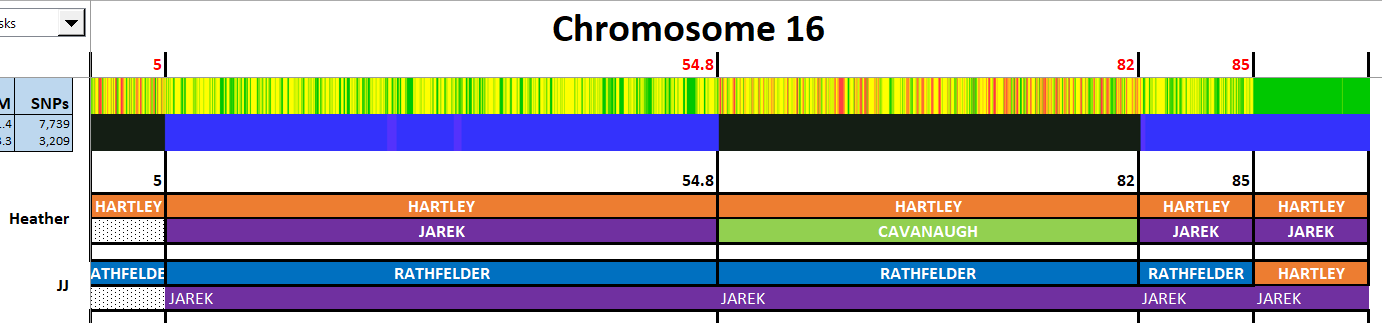
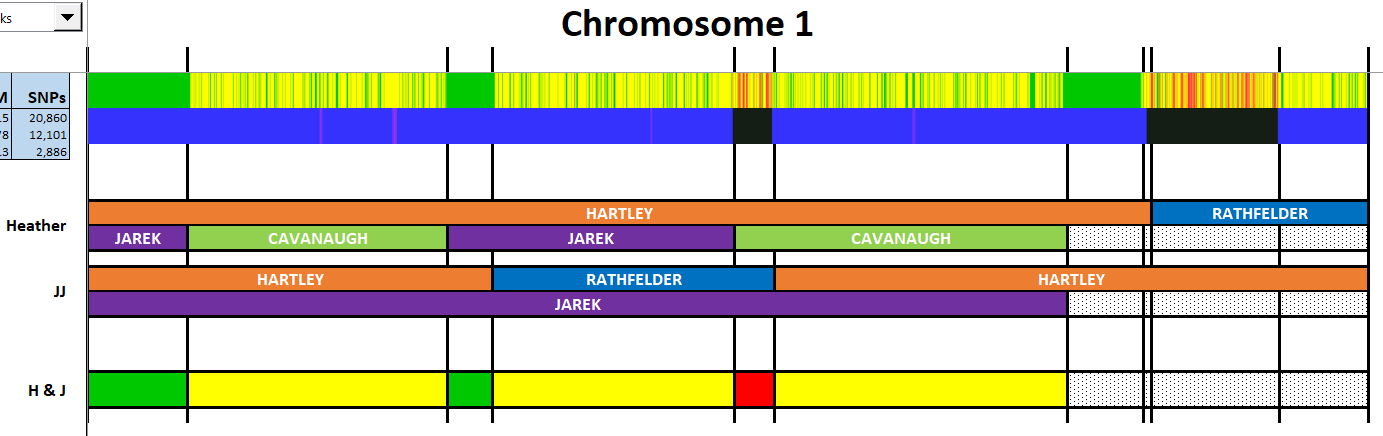
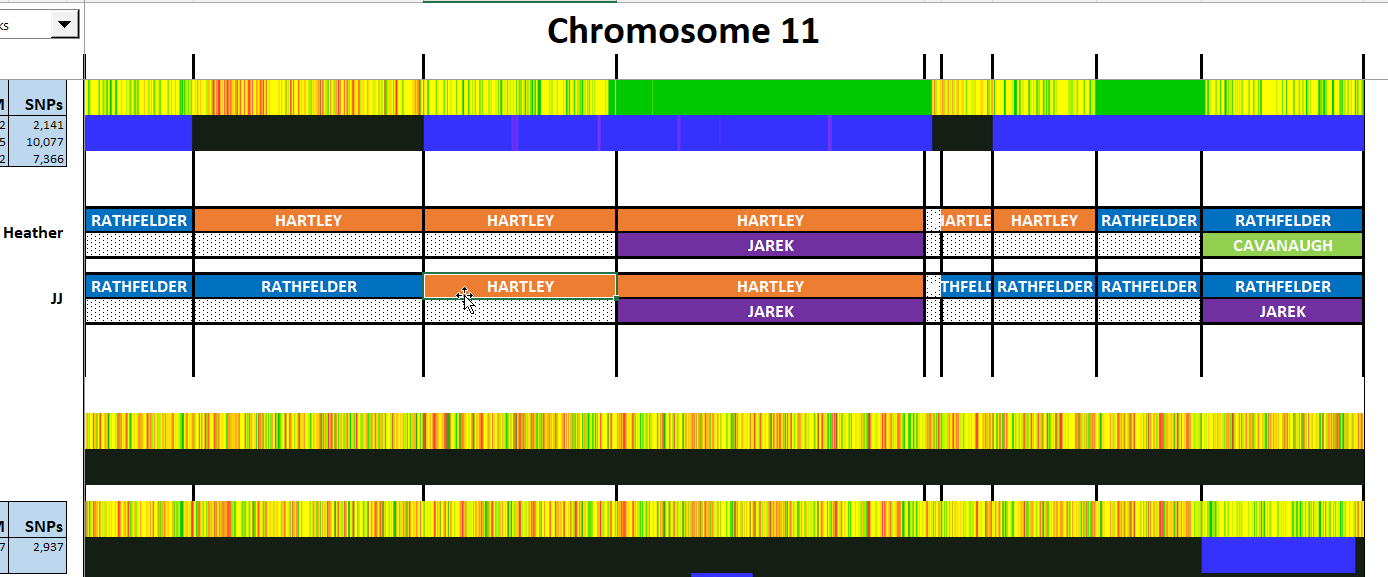
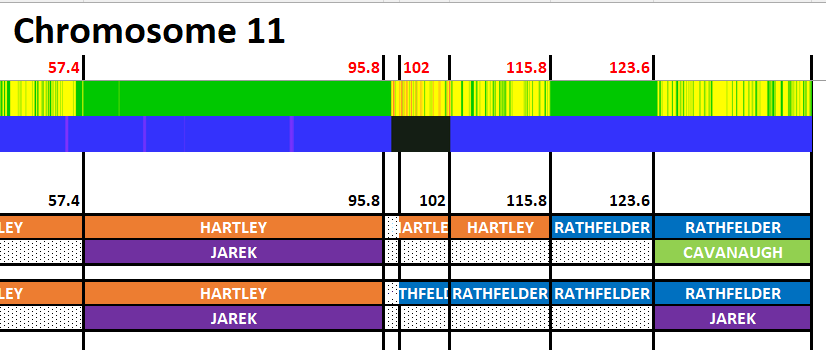
Here is my start at Chromosome 1:

I’m sure that there could be more hidden crossovers, but I started with what seemed to make sense. I already know that G2 is Rathfelder for my mom, Gladys. Where my children do not match my mom, I assume that they match my dad who I have as G1. There may be a way to go directly to a known grandparent, but I couldn’t figure that out. [I figured it out later by typing the surname into the cell.]
Robert on the Jarek Side
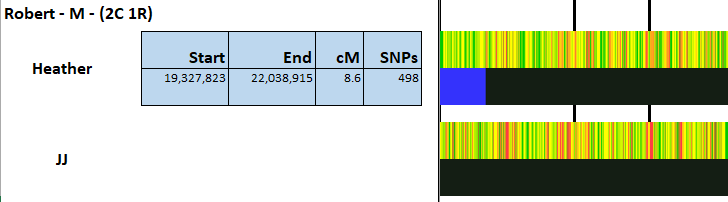
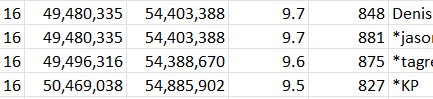
Robert matches JJ here:

As the match appears to go beyond the two crossovers, I will assume that three areas on JJ’s side are Jarek and the same three areas are Cavanaugh for Heather as she does not match Robert there. I’ll use G3 as Jarek and G4 as Cavanaugh. Unfortunately, that has caused a problem:

This mapping created a red region that has gone too far to the left. That means that either:
- The crossover should be further to the right, or
- Heather may have a small match with Robert in that large red section
If #1 is correct, JJ should be G4 (Cavanaugh) in the larger red section. If #2 is true, then Heather should be G3 (Jarek) in the large red section. I’ll check option #2 at Gedmatch. I can lower the threshold as far as 3 cM. I think I used to be able to go lower:

Now I need to look at the position numbers. Crossover #3 should be a 113M:

Heather has a match with Robert from 110 to 113M. That means that Option #2 is in play and Heather has a maternal crossover at that point.

Problem solved.
More Maternal Side Matches Needed
I added 4 more matches on Heather and JJ’s maternal side and deleted the Hartley cousin:

This added some information for Chromosome 1:

However, because I had not deleted the extra columns to the right of the phasing part, the new matches stretched way out. Mat2 shows a match with JJ and Heather in HIR which cannot be right. When I check where the first FIR goes to HIR it is just before 15M:

That is about where the match ends between Mat2 and Heather and JJ. Actually Heather’s match with Mat2 ends slightly further than JJ’s, so I’ll assume that Heather has more Jarek (G3) DNA there. However, I see that Robert has a match there also, which may be more accurate:

I guess the more matches, the better. I think that is as far as I can get with Chromosome 1. I just need to change the G’s to real grandparents.

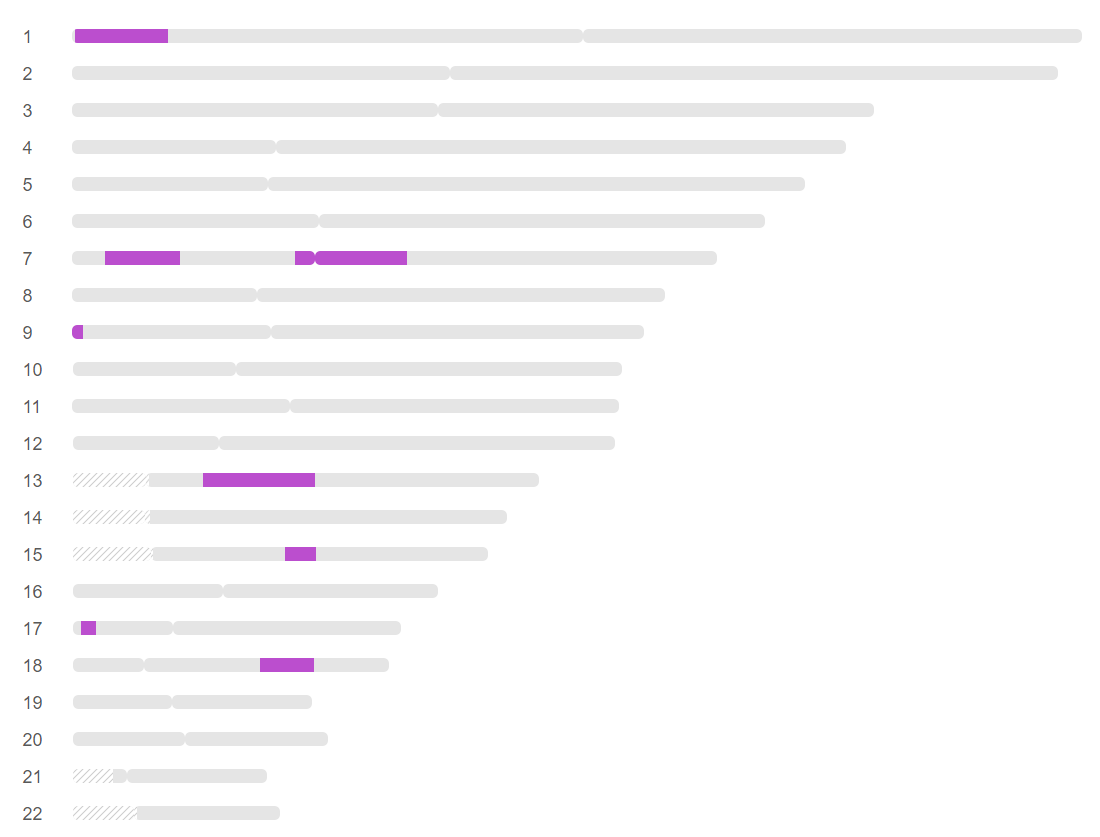
I have one more trick up my sleeve. By searching my old Blogs, I did find a Cavanaugh side match at FTDNA. I have used DNA Painter to represent the matches for Heather:

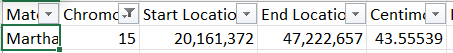
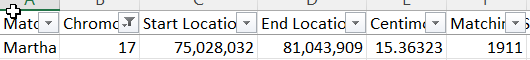
This shows that my guess for Heather’s first Cavanaugh segment was correct. I just need to figure out how to find JJ’s matches at FTDNA. By going through a bunch of ID’s, I was able to find JJ at FTDNA. I couldn’t find Marti, but found a Martha:

More importantly, here is Heather and Martha using the FTDNA Chromosome Browser:

I would say that Martha and Marti are the same person. As it turns out, the match does not help any further with Chromosme 1.
Chromosome 2

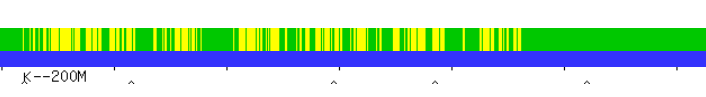
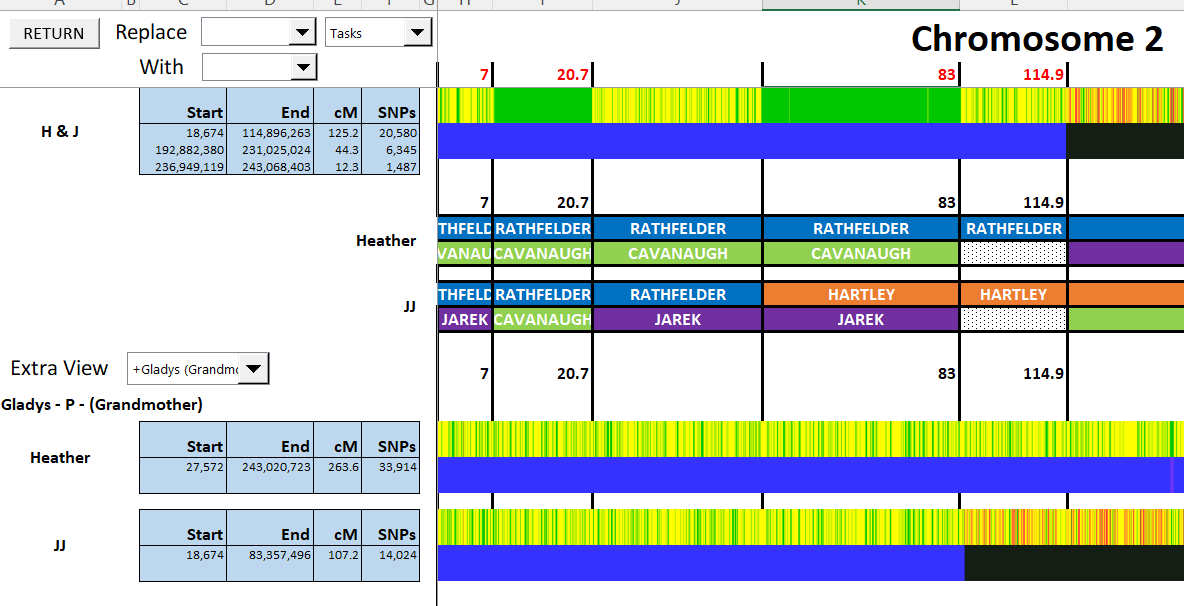
The bottom match is with my mother, so the Rathfelder side.
LeeAnne Match on the Jarek Side

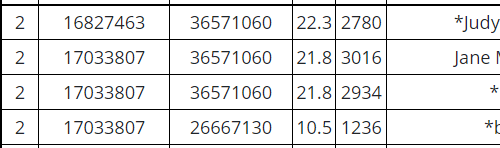
Here, I need to make an assumption. My assumption is that the right side of LeeAnne’s match with JJ and the right side of LeeAnne’s match with Heather represent crossovers. This is due to the fact that these end match positions are so close to the crossover positiions. There is a chance that these could be a coincidence, but I’ll take that chance.
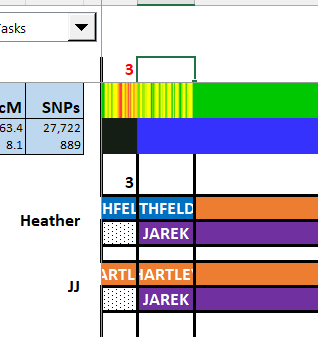
In additions, I see that Heather and JJ match Martha on the Cavanaugh side beginning at 7M. That is where JJ’s match with LeeAnne ends. That ices that crossover for me.
This seems to be as far as I get right now:

The fact that there are so few crossovers on the Paternal side, means that almost all the crossovers are on the maternal side. This makes this Chromosome more difficult to map.

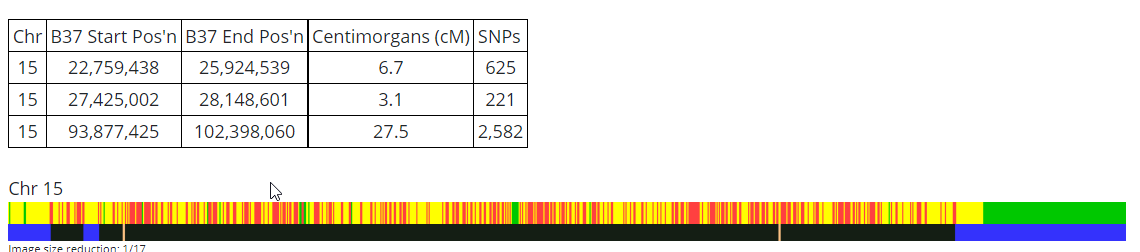
Chromosome 3
Just 21 more to go.

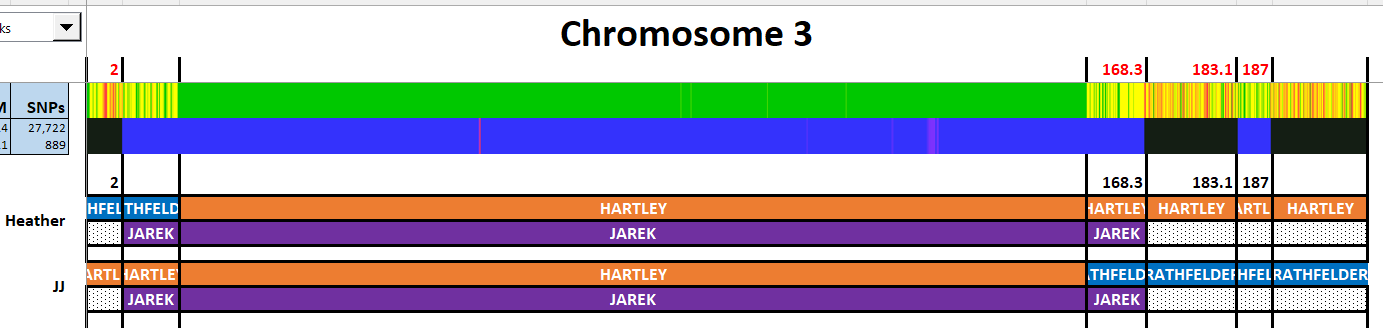
This looks fairly simple unless I am missing something. On the paternal side, it seems like there should be mostly Hartley. The blue match on the bottom represents Rathfelder. I see if I type in the names, I can skip the G1 and G2:

My next problem is that I don’t see any maternal side matches. I was ready to move on, but then checked MyHeritage. JJ matches Richard who has Polish ancestry:

Richard has an Anna Dziura. JJ has a Weronica Dziura:

These two could be sisters. Richard matches JJ on Chromosome 3:

All is not lost. Here is Richard and Heather:

They seem to match on Chromosome 3 in the same way.

Thanks to Robert, Chromosome 3 looks better for my children. This appears to be a mostly Hartley/Jarek Chromosome.
Chromosome 4
Here is what I get using my mother’s match and a Mat1 Jarek match:

Based on the the match of Mat1 with Heather and JJ, I would say that there is a maternal crossover to Cavanaugh to the left of the lowest Jarek on the map:

Also, note that there appear to be close crossovers between JJ and Heather indicated by the small black spaces in the comparison between JJ and Heather at the top of the chart. Heather shares DNA with Martha at the beginning of the Chromosome where JJ does not. This would indicate Cavanaugh DNA:

Also, JJ has a match with Martha here:

A Fully Phased Chromosome 4?
This is what I think is right:

However, one thing I would like to look at is the black break between 40 and 42M. I have no crossover there. I need to check that out more closely. Above, I note that JJ starts a match with Martha at 42M. That could be a crossover. Here is how Heather matches Martha:

Here is a corrected Chromosome 4:

The bottom red, yellow and green bar is helpful in checking to make sure the visual phasing matches with the comparison between Heather and JJ in the first bar above. Heather and JJ’s match with Martha was helpful in mapping Chromosome 4.
Chromosome 5
As usual, the paternal side is easy:

When I compare Robert’s matches with Heather and JJ, it points out a maternal crossover for JJ:


Here is Heather’s match with Martha on the Cavanaugh side:

JJ’s match goes further, so 160M is about the point of Heather’s crossover:

Next, I see that Mat3 has a match with JJ that doesn’t show with Heather:

This is at about 126 to 136M.

Going right on Heather’s bar, I can extend the matches:

This Chromosome is mostly completed except for the left side. There is a way to predict that side also. I just need to look at JJ or Heather’s matches to see which ones stop at the crossover and which ones do not. First, I need to find out where the crossover is. For that I go to the full resolution comparison between JJ and Heather at Gedmatch:

I’ll call that 32.5 M.
I have a list of Heather’s maternal matches that I downloaded in 2019:

The two maternal matches I have highlighted above appear to go through 32.5 M. However, I should also check with JJ’s results. I don’t see that JJ matches the same person. That makes me think that it is JJ who has the crossover at 32.5 M.

So, as I go along, I am remembering some of othe tricks of visual phasing. Here is the merged version:

Also, I have some position numbers as they are useful in mapping. Normally, there are not small isolated segments like Heather has on the right hand side (green Cavanaugh). This might be something to check.
Here is a maternal side match that Heather has:
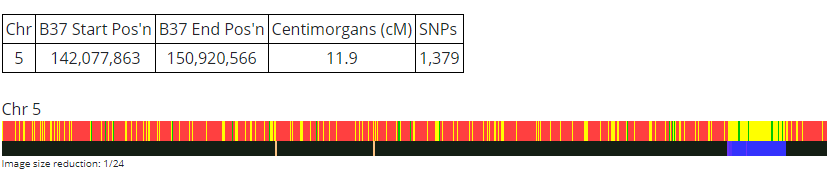
This appears to pass through 149.7. I don’t see that JJ has a match with this person.
A Corrected Chromosome 5 for Heather and JJ

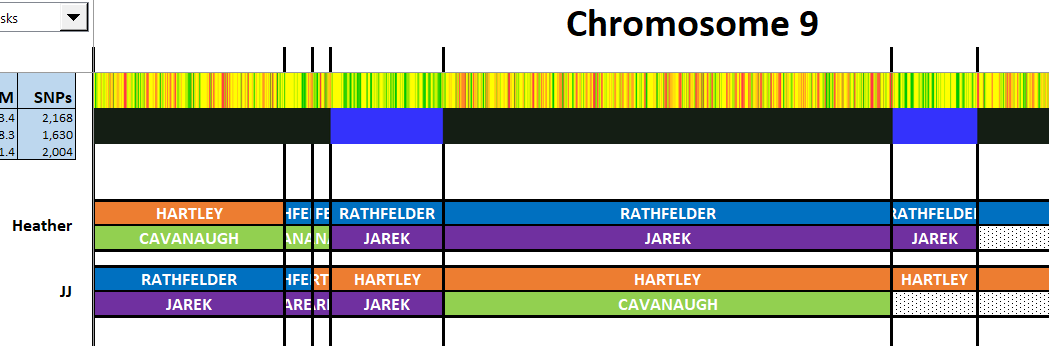
Chromosome 6
First I adjust the vertical lines and add in some of the position numbers:

Next, I check Heather and JJ’s match with their grandmother:

I see their matches do not line up well with the segment lines I have drawn. I suspect that the place where Heather and JJ both do not match with their grandmother should coincide with the green fully identical region between Heather and JJ. I’ll do a full resolution comparison between JJ and Heather to see where the green area is:

It starts at about 91.4M and:

ends at about 128.2M. These should correspond with these two match with Heather and JJ’s grandmother:

There is a bit of discrepancy between 126.6M and 128.2M.
That phases the paternal side for Heather and JJ:

However, I have no matches for Heather and JJ’s maternal side that were pre-loaded to the Fox Spreadsheet.
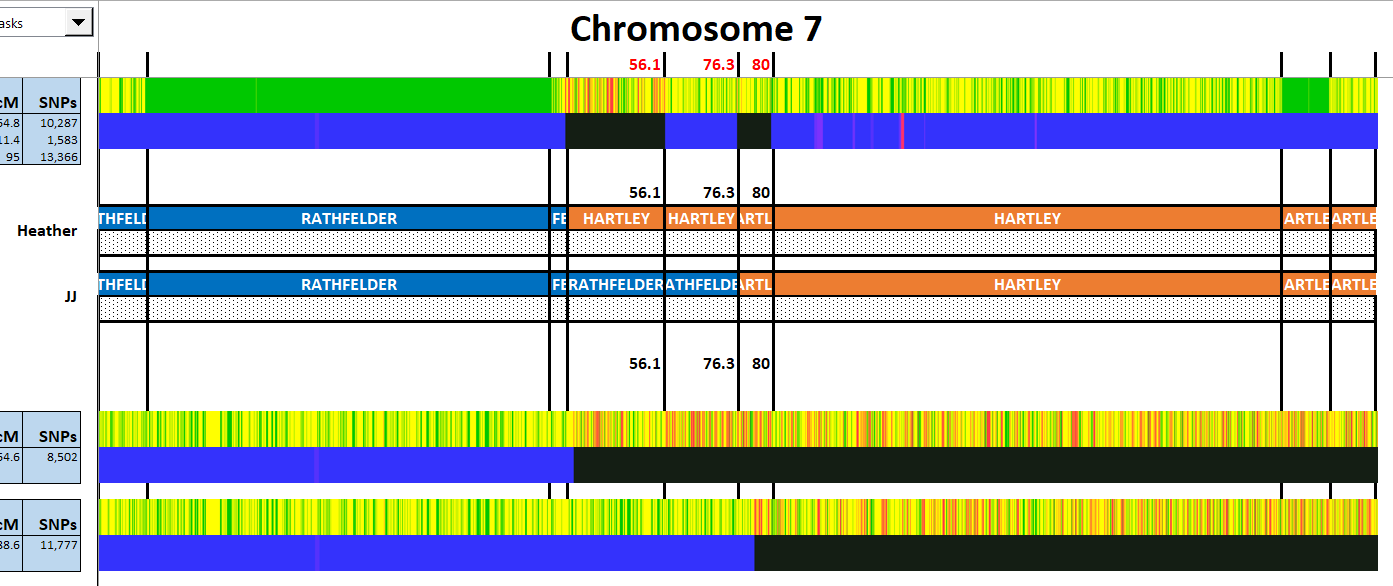
Let’s try Martha who matched at FTDNA on the Cavanaugh side;

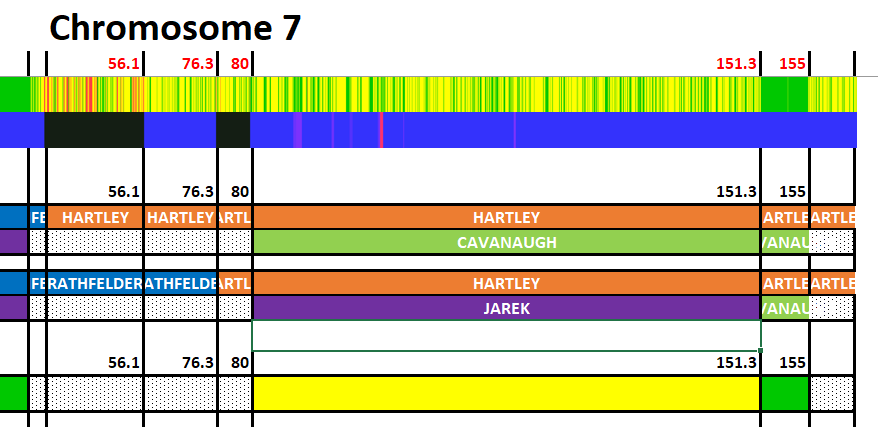
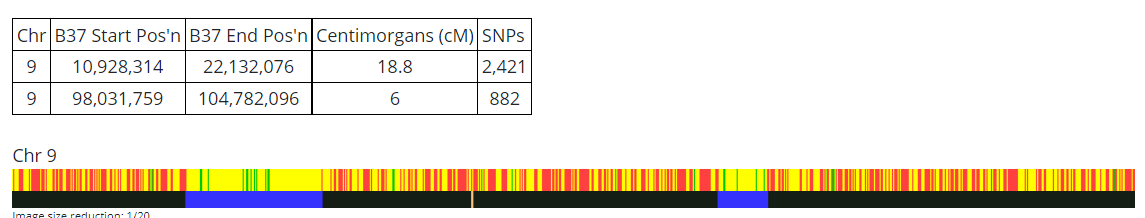
Martha has two matches with JJ. The first is roughly 7M to 42M. The second is:


The area above JJ’s green Cavanaugh must be Jarek for Heather as they don’t match at all in that region. The area above JJ’s second green Cavanaugh match must be Cavanaugh for Heather as this is an HIR or Half Identical Region. But first, let’s check on any match Heather has with Martha:

This points out a problem with my original mapping. Where JJ matchd from 7-42M, Heather matches from 4-14M. That means that I missed a segment here:

However, even that does not seem right. When I check at Gedmatch at a resolution of 3cM, I get this:

It’s odd how the comparison changes. [I explain my mistake further below.] Without any Jarek side matches, this is how Chromosome 6 is emerging:

At this point, I could look at Heather’s maternal match list to see where her likely crossovers are.

This 2019 list is a bit outdated. The last two matches are interesting. The last is with JJ and the next to the last goes through that match with JJ. That could indicate that JJ has a crossover at about 149. Here is JJ’s current match with that same person:

Here I added that crossover at 148.7 for JJ:

Using the same logic, I see this on Heather’s maternal match list:

Heather matches JJ from 72 to 135, but matches Sa from 68 to 92. That appears to give JJ a crosover at 72:

Unfortunately, this information is not helpful yet as I need to assign crossovers at 42.7M and 135M first.
Maternal Matches for JJ
I will check on JJ’s maternal matches to see if I can find some crossovers for Heather:
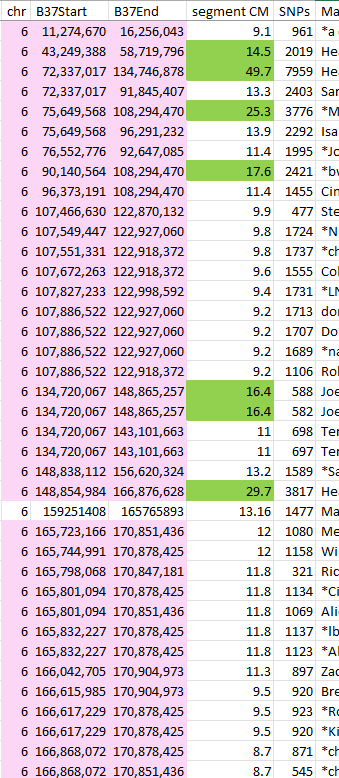
I didn’t realize I had this already and signed up for Tier I at Gedmatch. Oops.
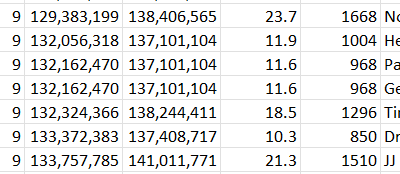
Here I see two matches JJ has starting at 134.7M:

This could indicate a crossover for him at that point. Now that I have Tier I, I will use a utility called segment search and use Heather’s maternally phased kit:

Here Heather has an important match with Jayne that is between 130 and 147M. I’m not sure why the match did not show up on Heather’s spreadsheet as it is 2399 days old. That clinches JJ’s crossover at 134.7:

I can move the Cavanaugh segments to the left as there are already paternal crossovers there. This gives JJ a small Jarek segment which we don’t like to see, but it seems to fit as of now.
That leaves crossovers needed to be assigned at 42.7 and about 72M.

Above, Heather has a match that goes through 72M, so JJ must have a crossover there.

Above, Heather’s match with Sara goes through 68.7 also. Plus JJ would not have such a small Jarek segment. Unfortunately, this has resulted in an extra unwanted crossover here:

Under my original configuration, this was not as much a problem, but when I added the crossover at 14, it is now a problem.
Lastly, I would like to know what is going on at position 167M:

JJ has a lot of matches going through 167, so I will give the crossover to Heather:


Here I tried taking the crossover out at 14, but the segment checker at the bottom was off. Part of the problem was that I forgot to give Heather a crossover at 68.7.

This solution appears to work out. I had made another mistake in that I had the spreadsheet scrolled over so that I couldn’t see the first HIR. Then I added a second crossover that I didn’t need at 14M.
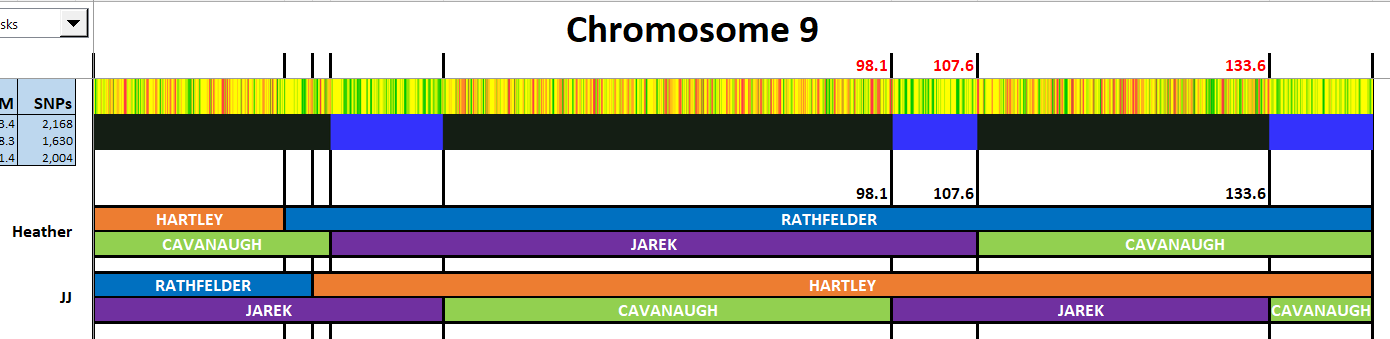
This version looks good, but may need some future adjusting:

There were fewer and smaller Jarek segments compared to Cavanaugh which could explain why there were not many Jarek matches on this Chromosome. It appears that this Chromosome could be a good one for looking for Cavanaugh side matches.
Summary and Conclusions
- I had some fun brushing up on my visual phasing skills
- Having JJ and Heather’s paternal grandmother tested made it easy to phase the paternal side
- Most matches on JJ and Heather’ maternal side are on the Jarek side
- There was one important maternal Cavanaugh side match at FTDNA
- It should be possible in most cases to find who gets the maternal side crossovers by looking at JJ and Heather’s maternal match list or by doing a Segment Search on Heather and JJ’s maternally phased kits at Gedmatch.
- I didn’t utilize this trick for the earlier chromosomes, so I will need to go back and do that at some point.





 .
.