In my last Blog, I mentioned that my brother Jon’s DNA test results came in this week. This happened in the middle of my attempt learn how to phase the raw DNA data for my 2 sisters and myself. I was phasing the data in what I can only assume is a traditional way. I say I assume, as I haven’t seen any other blogs on the process. The difference is that I am using MS Access which I hope will speed up the process. I should be able to get results for 23 chromosomes at a time instead of just one at a time.
The arrival of the new DNA results poses at least two problems:
- The previous 4 DNA data files were all in AncestryDNA version 1. Jon’s is in AncestryDNA2. While they are all Build 37, they look at somewhat different points on the chromosomes
- One of the difficult parts of the previous process was identifying and dealing with patterns of phased paternal and maternal bases. Those patterns were AAB, AAB, and ABB. With 4 siblings, there will be more patterns. However, the Whit Athey Paper I have been following does also look at 4 siblings.
AncestestryDNA Version 1 Vs. AncestryDNA Version 2
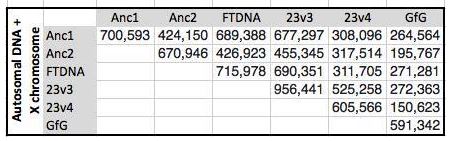
My understanding is that Ancestry changed the locations on the chromosomes that they were testing to get more into the medical area like 23andme. I don’t know if that is true. Here is a chart comparing the different atDNA tests:
I was doing well comparing Anc1 with Anc1 as I was looking at over 700,000 base pairs among 4 people. Once I compare Anc2 to Anc1, that is number is cut down quite a bit. That is about a 40% drop. My only other option, other than re-testing Jon, is to compare Jon to my mother’s FTDNA results. However, that will only pick up 2-3,000 SNPs, so I won’t bother.
Back to Square One with 4 Siblings: Homozygous Siblings
I need to find Jon’s equal base pairs and apply one to his ‘from dad’ column and one to his ‘from mom’ column. That is, after I add all Jon’s data to my database and add those columns. First I need to decide where to add Jon’s data. I could add it to the beginning of what I have already done or to the end. I’ll try adding it to the end, because I think that the work I did already is OK. I want to build on that. So rather than adding Jon’s DNA to the first step, I’ll add it to my table called tblMomBaseFromDadBase. This table has over 700,000 lines of bases for 4 people. Jon’s has 668,942 lines. Actually, when I remove “Chromosomes” 24-26, I will only have 666,531 lines.
Querying Jon into my latest table
Here I am adding Jon and the Mom from Dad Table to my query design:
Access thinks the ID that it added was important, but it really isn’t, so I need to take out that equal join. I really want the join to be at the rsid, but I don’t want an equal join. Why not? If I had an equal join, I would end up only with the positions that Jon has. I will lose 40% of the work that I have already done. Instead, I’ll use an unequal join.
I flipped the 2 tables in the query design area, so things are moving left to right. Then I choose a #2 join which is basically, an unequal join left to right.
Actually, I changed my mind. I have a better idea. I will just do the first 2 steps on Jon’s raw DNA and then join the results together. That is a third way that I hadn’t thought of. The point is, that there are many ways to do things in Access. There can be more than one way to get to where you want to be.
Back to Homozygous Siblings
First I copied Jon’s raw data into a table called tblJonHeterozygousSib. This is so I can use an update query to update the data in the new table and still have the original. Hold that idea. The better idea is to use a make table query. The reason that this is better is that it can take out the “chromosomes” I don’t want:
I took out the table I copied and I’ll make a better one with only Chromosomes 1-23. I hit the Run button and create a table with 666,000 lines:
Then in the above table, I inserted 2 rows: JonFromDad and JonFromMom. Now this table is ready to phase for any homozygous siblings. By the way, it looks like my Chr23 or X is homozygous, but it isn’t. Ancestry adds an extra base. I only really have one for my X Chromosome.
Finally time to query and phase
I go to Query Design in Access and choose the above table. This is a very simple Update Query design:
This says if Jon’s allele1 is the same as his allele 2, put allele 2 as his base from mom and as his base from dad. I hit the run button for the update and get the dire warning that I’m updating a lot of information, I can never change it back. Then I get a message that I’m updating 478,000+ rows. That is good. Those are the number of Jon’s homozygous bases – quite a few. I’d say over two thirds.
I’m not looking for crazy results and didn’t get any.
Homozygous Mom Query
I’ll copy my previous table into one to update. Then I need to add Jon’s base from mom where mom is homozygous. Easy peasy. I think this is all I need.
Actually, I did think of an issue. I have an equal join. That means I won’t be using the homozygous bases that mom tested for in the old AncestryDNA test that aren’t in the new AncestryDNA test list. My guess is that is interesting information but perhaps not very useful. It also occurs to me that in the spots where Jon doesn’t match up with my siblings, I will still have the 3 letter pattern work that I had done previously.
The query above says if Mom allele 1 = 2, then put that 2 allele in Jon’s from Mom base slot. I hit Run and pasted 277,000 rows of bases.
This query will be a little more difficult to check. I have to create a query linking my mom’s DNA results to this table. I did that and see one problem already.
The first problem is that ID 126 didn’t show up. That means that rs3819001 that Jon has is not in my mom’s raw DNA. I don’t want to have data for Jon that looks like it can be updated, but it can’t.
I think I can fix this.
Updated Table Query
A few steps ago, I ran a Table Query to get just Chromosomes 1-23 into Jon’s Table. I need to upgrade that query so that I am only including the locations (rsid’s) that are common to both my mother and Jon. I do this using an equal join on the rsid Field:
This time, my table for Jon only has the rsid’s that my mom has.
Also my Chromosome formula was off, so I had to fix it. Also note that I have about the number of rows as per my Anc1 vs. Anc2 table earlier in the Blog. I then re-added the Jon from Dad and Mom columns into the new and improved table. Then I reran the update query which told me I was about to update 284,000+ rows.
This worked as well as last time, but this time I have the fewer rows I was trying to get.
Re-Run the update query for homozygous mom for jon
I double clicked on my old update query. The message said I was updating 277,000 rows or so. Now I’ll re-check my work. If there is no ID 126, I’ll be happy. Well it is still there, because I forgot to copy the previous homozygous sibling table into the homozygous mom table. After re-re-running the update, I got the desired results:
And there you [don’t] have it: no ID 126. Here is my mom’s raw file compared to Jon’s updated table.
Jon gets a G from mom at ID 128 even though Jon is AG, because mom is GG. Now I’m talking DNA.
Merge Jon’s New Table with His 3 Siblings’ Tables
This is the point where I put everything together. I will try to use the Make Table Query for this one again. So I’ll put my newest Jon table together with my newest sibling table.
This shows the left to right arrow join. I’ll want the larger file plus everything equal in the smaller file. Come to think of it, this Create Table Query would have fixed the earlier problem I had. I guess I was too careful! The other issue is that the ID in the 1st table won’t be the ID in the second table. I could keep the second ID, but I would have to rename it as Jon ID or Anc2ID.
Here I rename Jon’s IDs as JonID. I may not need it, but if I do need it I will have it. I guess MS Access wasn’t happy with my idea:
OK, I took out the JonID and hit Run. Microsoft tells me about my new 700,000 row table.
Back to the Dad Patterns
Now that all the family is together I want to look at Dad Patterns, because I know that I will be updating those. Here is the first query I tried on my new Table of 4Sibs.
This is looking for filled in Dad bases where Sharon’s base is not the same as Joel’s. That query gives me an ABAA pattern:
Also ABBB:
Here’s ABBA:
It looks like ABAB is a possibility also. That means the following are possible:
- AAAB
- AABA
- AABB
- ABAA
- ABAB
- ABBA
- ABBB
So if I chose Joel’s Base not equal to Sharon and then Joel’s base equal to Sharon would I have every combination? It looks like I need this combination to cover all possibilities:
- Joel <> Sharon OR
- Joel<>Heidi OR
- Sharion<>Heidi OR
- Heidi<>Jon OR
- Jon<>Joel OR
- Jon<>Sharon OR
Which in Access looks like:
But Wait, I Forgot Principle 3 for Jon
Principle 3 says where Jon is heterozygous and he knows where he got his maternal base, the other base goes into his From Dad column. Looking back at my old queries, I see this is a 2 step query. I’m tempted to try this in one step, but I think this got me in trouble before, so I’ll go with the simpler query. Simpler queries are usually better in MS Access.
This says where Jon is missing a phased allele from Dad and he has an allele that doesn’t equal the one he got from mom (making Jon heterozygous here) put that allele into Jon’s From Dad spot. I tried the query and only got 37 results. The problem is, I should have said ‘Is Null’ in the JonFromDad Criteria:
This time I get 35,000 updates, so that is right. I then change the allele1’s to allele2’s above and get 33,000 updates to tbl4Sibs. I ran a quick query on the 4Sibs Table to get just Jons heterozygous results:
In the first line, Jon had allele1 as T which was different from the allele from Mom of G, so Jon’s T got put into the From Dad spot. At ID 41, Jon’s allele2 of G is from Dad because he had an A from Mom. When parent and child are heterozygous, the From Parent location remains blank.
Now I have Jon with 3 Principals: Homozygous Jon, Homozygous Mom and Heterozygous Jon.
Back to Dad Patterns
I have the old Dad Patterns for 3 siblings. Now I need to See what the 4 sibling Dad Patterns would be and add Jon’s Start and Stop Locations for his new Dad Pattern Areas. I need to combine that with the 3Sibs Table.
My first query was wrong and gave bad results. The reason is that the ID for 4Sibs was from the raw data. The ID for the Dad Pattern Table just numbered the amount of Dad patterns. I needed to join the ID in the first table to the start and stop locations in the second table. I ended up doing 2 queries: one for the start position and one for the stop as I needed both. This query gives the stop position of a pattern.
I took both those queries and put them into an Excel Spreadsheet.
I added a new column called Dad4Pattern. In the first row, the new pattern was AAA by chance. However, in the second row which is the Stop or End of the first Dad Pattern, it is obvious that the ABA Dad Pattern goes to an ABAA Pattern. I didn’t think that there would be many AAAA Patterns as that means that all siblings match the same Paternal grandparent. This is the only AAA pattern that I had noted so far as I wasn’t looking for them yet. Still, I will need to go back and verify that these Start and Stop AAAA’s were not by chance. Finally, on the last line, it is clear that the Dad Pattern goes from AAB to AABB with Jon added.
Next I chose all the cells where Jon had a base from Dad and performed a Concatenate operation to write the pattern.
This gave me the CCCC that I wanted to check. Next, I wrote a formula to put the Dad bases together in a new column and wrote down the Dad Patterns that I had.
A few notes:
- Out of the 66 three sibling patterns that I had, I was able to find all but 5 new four sibling Dad Patterns. See the yellow above for two of the missing 4 sibling dad patterns.
- The missing 4 sibling dad patterns should be easy to find by scrolling through the 4Sib Table
- I noticed that there were no AAAB patterns. That is because in my previous search, I was not looking for AAA patterns. So now, I don’t have any AAAB patterns. I will have to find these in my new search.
- AAAB is the situation where I match the same paternal grandparent as my 2 sisters, but Jon matches the other paternal grandparent.
Filling in more dad patterns
To fill in the yellow areas, I made a query in Access based on the 4Sibs Table. This looked at every case where Jon had a base from Dad. Searching around the ID 6604 and after, I found this pattern:
ABBB
Then I checked near the end of the old 3 sibling pattern which is at ID 19806.
At ID 19827 we see an ABAB Pattern, so I enter that Pattern in my spreadsheet:
For the start of the new ABAB pattern, I used the old ABA location as that was more precise. The next interesting thing happens at Chromosome 2:
Here I have a problem in my spreadsheet. For some reason, the Start of the last pattern of Chromosome 2 ends at Chromsome 3, which is not right. My previous spreadsheet was better than that. From the ashes I will re-build.
I note that at ID 108798, my 4 Sib Spreadsheet goes to an ABAB Pattern. At the end of Chromosome 2, I see an AAAB Pattern. That was the one I wouldn’t have had from the 3 sibling pattern as I wasn’t checking on AAA’s.
I added new rows for the patterns ABAB and AAAB:
The most important thing here is the ID, the pattern, the Start and Stop. Here is the new change area from ABAB to AAAB:
There are a few SNPs between the ABAB Stop and the AAAB Start that are a little unclear.
Finding Jon’s Patterns
Now I’ll check Jon’s Patterns. I’m looking for any changes in patterns as these should be important as crossovers later. I will need to assign the crossovers to each sibling’s Chromosome Map.
Good Old Triple A – B Pattern and all the others
AAAB is where Jon has a different paternal grandparent than his 3 tested siblings and the 3 siblings have the same paternal grandparent.
My query says that Jon has to be different from each sibling. I run that and insert the appropriate Start and Stop point for the AAAB in my spreadsheet.
I do the same for AABA which I can find using a similar query under Heidi’s criteria:
I ended up going to a clean spreadsheet. It was too messy combining the 4 sibling results with the old 3 sibling results.
Here I have the ID, the Chromosome, the pattern and the Start and Stop. The yellow marks a one SNP pattern. It appears that there should be 3 types of patterns:
- One where one sibling matches none of the others. That is what I have above: AAAB, ABAA, AABA and BAAA
- One where 2 pairs of siblings match each other: AABB or ABBA. I’m not sure what else there could be. I looked above and saw one other: ABAB
- One where all the siblings match each other: AAAA
That makes 7 or 8 patterns, depending on whether AAAA is considered a pattern.
Two Pairs of siblings match each other patterns
Here is the Access query for AABB
At first I was missing the criteria under SharonFromDad and that gave me AAAA combinations also. The result of the query looks like this:
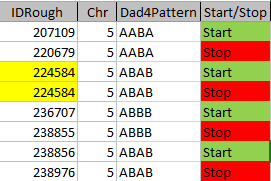
Here Joel matches Sharon and Heidi matches Jon but on a different base. After I was finished putting in Starts and Stops for each Pattern, I then sorted my spreadsheet by ID. This brings up some issues that need looking at:
Where there are 2 Starts or Stops in a row, there is a need to check what is going on. The ones around the yellow positions may not be a problem as I’ll likely be taking those single positions out. However, at the end of Chromosome, there are 2 starts and 2 stops together. I need to go to ID 236707 and see what is before that point. It apears that there is an AAAA pattern before that point and that the ABAB at 224584 is a single point. That fixes half of the problem. Then I go to ID 238976 to see why I have a Stop there for ABAB.
I had missed the Start for the ABAB right after the stop of the ABBA pattern, so I added it in. The repaired spreadsheet looks like this.
An application
Now that I have the change between ABBB and ABAB described, let’s look at what it means. Here is a different look at that location:
When the pattern changes from ABBB to ABAB, what has changed is the third B changes to an A. Heidi is in that location. So that says at the above position of Chromosome 5, Heidi has a paternal crossover. I thought it would be good to check my work against the work of M MacNeill. To do that, I used the NCBI Remap website to change my Build 37 results to Build 36:
This would be the start of Heidi’s new segment. Here is what MacNeill had:
I got it right again. That is 2 for 2. Actually, the first time I tried, I was comparing the wrong Chromosomes. Rookie mistake. Here is M MacNeill’s map for Heidi on Chromosome 5:
Perhaps it is difficult to see, but the point I am looking at is the little lighter red segment at the far right of Chromosome 5. Perhaps that is why I missed it the first time as it is so small.
Another Aside is that this was a very difficult Chromosome to decipher using visual methods. This was one of my attempts to figure out the crossovers visually for 3 siblings.
I had missed the last crossover as it is so small and difficult to see. In my defense, I should note that M MacNeill did mention that the end of this Chromosome was difficult to decipher.
Taking Out the X
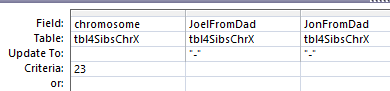
I’ve realized that I’ve generated some bases for the X I got from Dad. Of course, I didn’t really, so I’m taking out any bases there for me and my brother Jon. I’ll use this update query:
I was worried that I’d mess something up, so I created a new table called 4SibsChrX. My query put dashes in the spots where I couldn’t have an X base from Dad:
This looks like a good place to end Part 4. It appears that there should be many chances to quality check my work and that the process is progressing. Getting Jon’s new DNA set me back a bit, but the results should be better than what I’d see with 3 siblings.