Right now I am looking at my sister Heidi’s DNA matches with Common Ancestors.
These 7 matches could keep me busy for a while. Three maternal. Three paternal and one unassigned. These are all listed under distant relatives.
Heidi and Sonja:
Sonja has her Hield ancestor as being from Connecticut, so no obvious connection there. I need to know if Sarah married a HIeld and moved to Connecticut. I do know that Sarah was living in Sheffiled in 1851:
Sarah’s father had died young and her mother was a beerhouse keeper to make ends meet. Here is Sarah Hield in Hartford, Connecticut in 1860:
We can see that Sarah’s daughter Sarah was born in Connecticut, so that puts the move from England between 1855 and 1858. Sonja has that Esther was born in 1861 so that explains why she does not appear on the 1860 Census.
Here is the possible marriage:
Unfortunately, the father’s name were “dead” which is not very helpful. However, I do know that ‘my’ Sarah’s father was dead. Here is PIttlsmoor:
Sonja’s DNA Connection: Shared Matches
Here are some of Sonja and Heidi’s shared matches:
Melinda is my maternal 1st cousin’s daughter. Carolyn is my mother’s second cousin on the Nicholson side. The other matches seem to be related on the Nicholson side or Clayton side. The DNA indicates that the Common Ancestor clue from Ancestry is probably right. This gives encouragement to continue along the lines of the Common Ancestor match.
Nicholson ThruLines
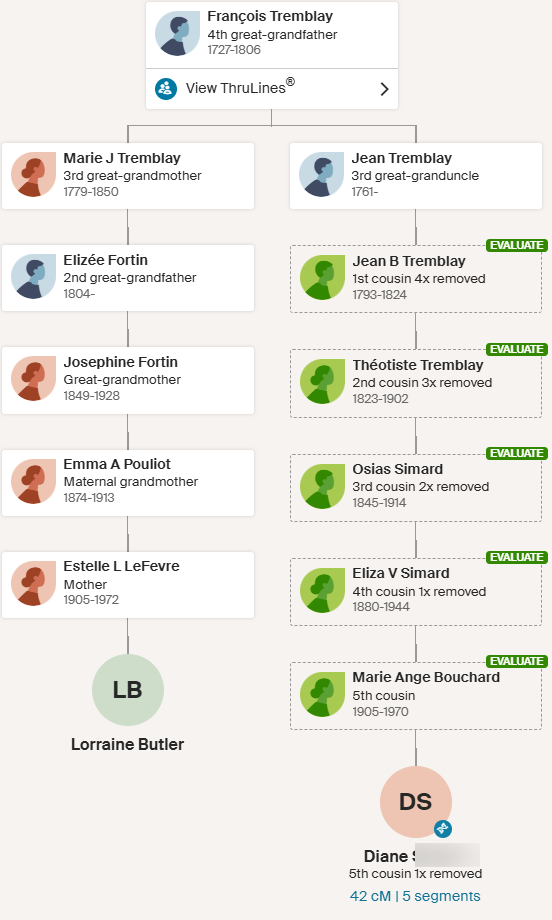
Here are Heidi’s ThruLines:
Back to the Genealogy
The Hield family seem to move around a bit. The first son, John William, was born in the Wicker, Sheffleld:
The name John Willam also gives circumstantial evidence to relation to the Nicholson family:
William was Sarah’s younger brother. John was her grandfather. We are not sure of Thomas Hield’s ancestry, but Sarah’s second son was named Thomas. As a guess, Esther could have been Thomas’ mother:
Let’s look at the proposed timing.
- Sarah Ann marries Thomas Hield in 1852
- They have children and move to Hartford, CT around 1856 or 1857
- Sarah’s younger brother William arrives in the US about 1868 with his family and settles in Philadelphia.
I’ll add Sonja to my tree as floating tree and likely connect her later.
Sonja’s great-grandfather was an interior decorator in 1920 in West Hartford, CT.
Here is the family in 1900. There were a lot of Russels:
This Russel was a stock broker. However, at this point, it is Esther that I am interested in. From the Census, it appears that Esther married about 1880.
Here is ab obituary from August 11, 1928:
Recall above that Esther had a brother named Thomas. Circumstantial evidence again. This leads us back to Brooklyn:
Sarah A HIeld is 39 in 1870 which means she was born about 1861.
Here is the Sarah Ann in my tree:
I’ll say that is close enough for a match. I just need to merge the two trees. Here is part of my new tree:
My ancestor William Nicholson was about 5 years younger than Sarah Ann. I wonder if William and Sarah Ann ever connected in the US.
Summary and Conclusions
- I started at looking at my sister Heidi’s unviewed Common Ancestors at Ancestry
- Heidi’s first match with possible common ancestors was Sonja
- Sonja’s tree went back to Connecticut with her ancestor Esther Hield
- Ancestry suggested that Esther Hield’s mother was Sarah Ann Nicholson, the sister of one of my Nicholson ancestors
- Based on Shared DNA matches between Heidi and Sonja, as well as genealogical clues, the match appeared to be right
- It would still be nice to find the smoking gun genealogical clear evidence, but the inferred evidence from the DNA and genealogy was enough for me to agree that Ancestry’s proposed common ancestors were correct.
- That leaves 6 other proposed comon ancestors that Heidi has at Ancestry to investigate




























































































































































































































































 .
.
















