In this Blog, I would like to look at my raw DNA data. Those are the A’s, T’s, G’s and C’s. I have tested at AncestryDNA as has my mom and 2 sisters, so I will use those results. Whit Athey has a paper that describes how to phase your DNA when the DNA from one parent is missing:
Downloading AncestryDNA: Getting Rid of Zeros
Many people have downloaded raw data to upload to gedmatch.com. Ancestry raw data is in text form. Access gets along with Excel well, so first I import the AncestryDNA text data into Excel. Perhaps if you are curious, you have taken a look at your raw data to see what it looks like. Unfortunately, it takes a while to open up such a large file. Here is what a few lines of my AncestryDNA text file look like:
It is important to note in the information above that Ancestry uses Build 37. That means that these results need to be converted to compare to Build 36. For example, Gedmatch uses Build 36. I remove the information above the column titles and bring it into Excel. However, I put my name on the top of the last 2 columns because eventually there will be columns for 4 people’s results (mine, my mom’s and my 2 sisters’). I will need to distinguish between each person’s alleles. It is important to note that when importing this text file to Excel, Excel retains the file as text. This is probably such a file as note that the no-calls have been changed to zeros. To save the file as an Excel file, you must specifically do that step.
Here is a file with the no-calls as blanks, like I want them, and with my name at the top and the verbiage removed:
Here is the file in Excel. I have used the search and replace in the last 2 columns. I want blanks for no-calls and not zeros which Excel likes to add.
Using Access
At this point, I had to switch to my laptop as I don’t have Access on my desk top. I open up Access and name a new database. I go to External Data and choose the Excel icon with the arrow pointing up to import my 4 Excel Files of Raw DNA for Mom, my 2 sisters and me.
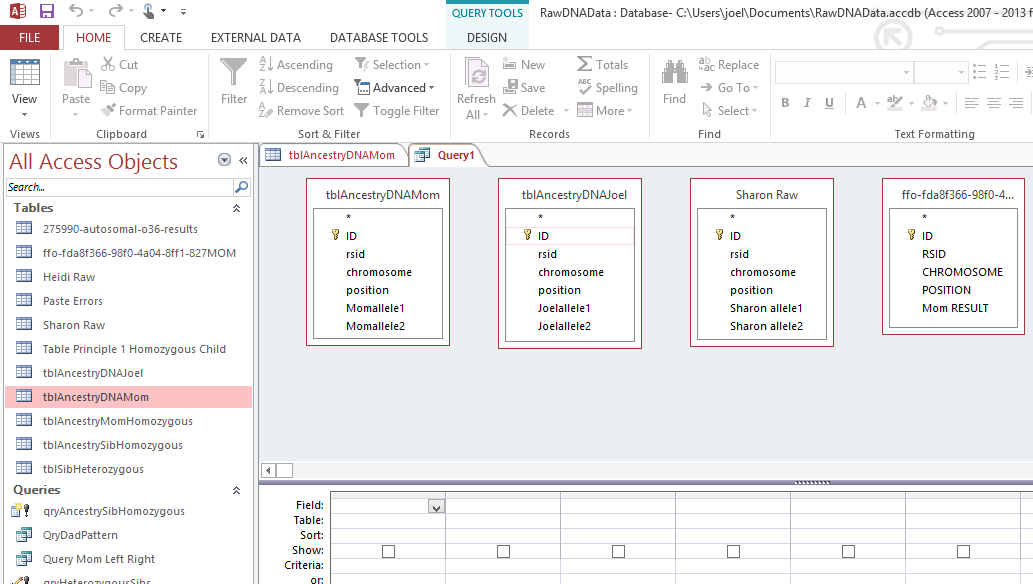
Next under Create, I choose Query Design. I choose the 4 Excel files that I have imported to Excel.
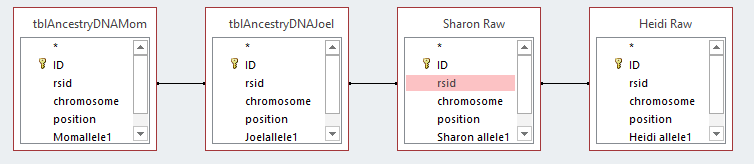
I should note that when I imported the Excel files, that Access creates a unique ID for each row. I let Access do that. It has set that ID as a key identifier. I could have used the rsid as a key that is somewhat as a unique constant. Next I will connect each table by the rsid’s with something called an equal join. That is the dark line I added between the rsid Field for each persons DNA data.
This means return the results when the rsid is the same for each file. Note the last table ( 2 images above) was wrong, so I took that out and added my sister Heidi’s Raw data table on the right. It is important to get the initial importing right and in the right format as this will save a lot of time later. Here is the form that I want the data in:
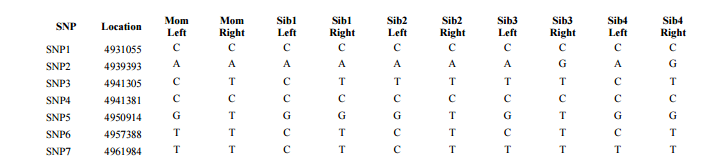
This is a portion of Table 1 from the Whit Athey Paper. The difference is that Whit only had part of Chromosome 16. I will have all Chromosomes at once. In my Access query I choose the Excel Titles as Fields. I need the rsid, chromosome and chromosome position only once. Then I add the 2 alleles for each person. FTDNA uses right and left alleles. AncestryDNA uses allele 1 and 2. They are the same undifferentiated alleles.
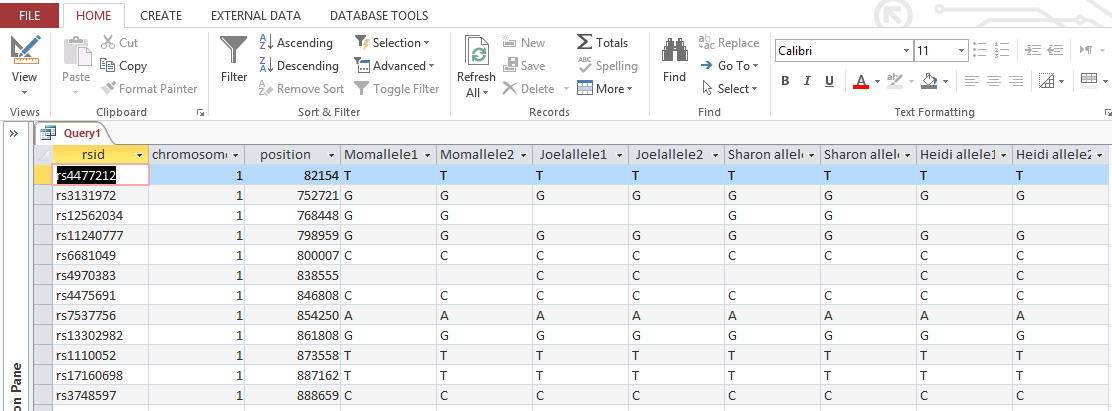
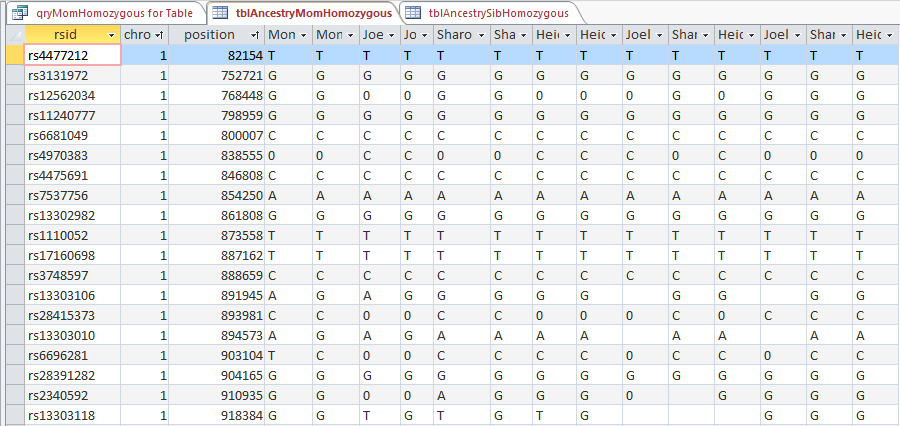
When I run the view the query results, I get this:
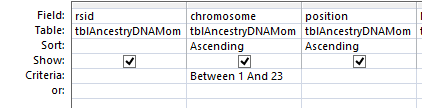
So with one push of the button, I have all the raw results of 4 people in my family in one area. I actually have more information than I need. AncestryDNA includes chromosome 24 and 25 which is YDNA and mitochondrial information that I don’t care about here. This is easily filtered out in the criteria section of the design view. I choose ‘Between 1 and 23’ there. That gives me each chromosome between and including 1 and 23.
Now I am down from roughly 701,000 lines of data to the 700,000 lines that I want. It is important to save these results as a Table in Access as we will be using that Table to make more tables. Also save the query. Even though I say to do this, I didn’t. but just saved the results under the next step.
Whit Athey’s Principle 1
This Principle is simple and straightforward. It says that if you have two letters the same in your results, one of those came from one parent and one came from the other. In line 1 of my results above I have TT. All my siblings have this result also. My mother is already shown as TT as she was tested. My father who was not tested must have had a T which he gave to me and my 2 sisters. Here is Table 2 from Athey showing the next set of data that we need to produce the AncestryDNA raw data. Ancestry didn’t tell us which side each of our bases came from, so we will figure that out.
I have only 3 siblings that I’m looking at right now, so I need 6 more ‘Fields’ in my database. There are a few ways to do this in Access. Here is one way that I did it.
Athey Principal 1 in Access: Homozygous Siblings
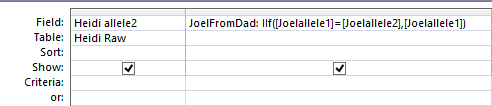
Homozygous is just a fancy term for my TT result found in the 1st position tested of my 1st Chrmosome. I created 6 more fields. These are to show what allele (letter) I got from my dad and my mom when I had a TT or other such homozygous results. Here is what the first field out of six that I added looks like on the Access Query Screen.
JoelFromDad is the first new field name. After the semicolon is the criteria. In English is says that if my allele1 is the same as my allele2, then put my allele1 in as the result I got from my dad. I used the same reasoning for a field called JoelFromMom and in similar fields for my two sisters. I viewed the results to make sure they made sense. I chose Make Table as I want the results in a Table to use later.
This will make sure that the chromosomes and positions within the chromosomes stay in the correct order. Otherwise, Access may try to sort by the first field which is the rsid.
Principle 2 in Access: Homozygous Parent
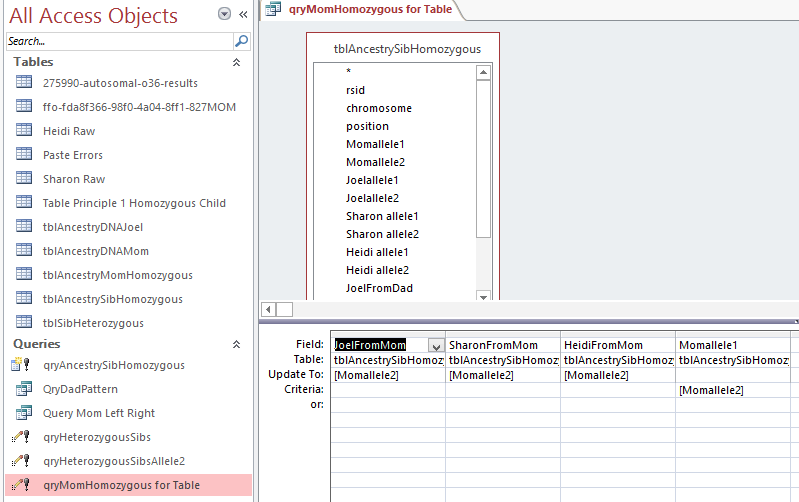
In my case, the homozygous parent is my mom. I spilled the beans already by my mistake above. In Line 3 above, my mom is GG. That means she had no other choice than but to contribute one of those G’s to each of her children at that location on Chrmomosome 1. Now I will put that Principle into Access language. For this portion I will use an Update Table. An Update Table will add new information to an existing Table. In this case, I added it to my tblAncestrySibHomozygous Table. That is why it showed the results already above. Here is what the Update Query looks like in design:
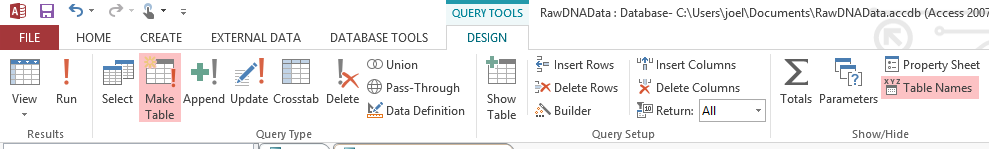
Here I have the tblAncestrySibHomozygous Table which I reran (or un-updated). This query says for the criteria where Momallel1 equals Momallele2, update the JoelFromMom, etc Fields with the Momallele2 value. Obviously I could have chosen either of her alleles to update the fields as they are the same. In the bottom left of the image above there is a pink highlighted query called qryMomHomozygous for Table. That is this update query. the ! means that it is going to create something. I assume that the little symbol to the left of the ! means that it is an update query. I ran the query and then created a new table with the results called tableAncestryMomHomozygous. Again, what I had forgotten was that by running this query, I also updated tblSibAncestryHomozygous. It’s always good to do quality checks – especially when you are dealing with over 700,000 rows of results at one time.
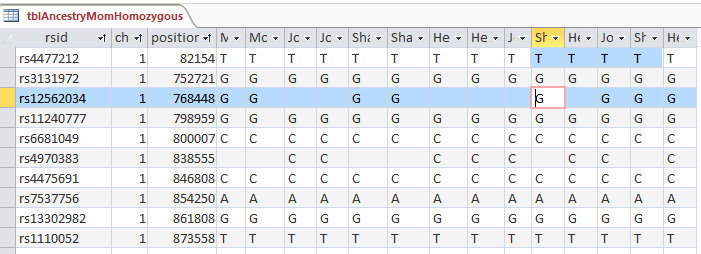
I did the update and got a warning from Access that I was updating over 400,000 rows. And that action cannot be reversed. Here is my old tblAncestryMomHomozygous to show the zeros that I didn’t like:
Principle 3: Heterozygous Child
I’ll copy the Athey Principle as he stated it as it is slightly more complicated than the previous two:
Principle 3 — A final phasing principle is almost trivial, but it is normally not useful because there is usually no way to satisfy its conditions: If a child is heterozygous at a particular SNP, and if it is possible to determine which parent contributed one of the bases, then the other parent necessarily contributed the other (or alternate) base. This principle will be very useful in the present approach.
How to put Principle 3 into access?
Here is an example of heterozygous children alleles where the mother’s contributing base is known:
We know that each sibling got a G from mom as she only has G’s at this location. All the siblings have TG for their raw results, which means the T must have come from dad. I can go through over 700,000 lines and apply that rule or try to use the Access Update Query to produce the same results. This time I copied the tblAncestryMomHomozygous to a table called tblSibHeterozygous before I did the update to maintain the integrity of the older table. In the Update Query, I combined 2 steps. First I set a criteria that there has to be something in the JoelFromMom for this to work. So I said that JoelFromMom is not Null. Next, my allele1 is T. I want this to go into my JoelFromDad spot. If this T doesn’t equal the G I got from my mom, I am already heterozygous, so I don’t need an extra query for that. [That was the step that I didn’t need.]
Here is what I have for an Update Query:
However, note that instead of looking at allele1 here I chose allele2. I am thinking that this will be a 2 step process for each allele. This query is updating over 70,000 rows, or a little over 10% of all the data. I’m trying to show that this Update Query did not address my example above which had to do with allele1 and it didn’t:
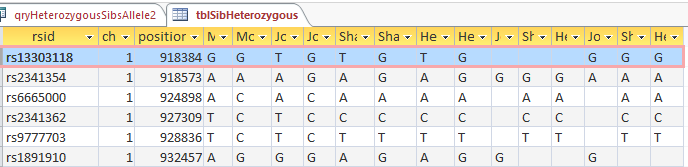
The first line was my example. The 3 blanks in the first line are the bases from Dad that were not produced from the query as expected. However, it did work for the second line. In that case, my allele2 was not equal to the allele I got from my mom, so it inserted that allele (G) as the allele I got from my Dad. Next I’ll copy the query: qryHeterozygousSibsAllele2 and rename it as qryHeterozygousSibsAllele1. Then I changed 6 of the allele2’s to allele1’s. This is to cover my original example where allele 1 wasn’t the same as the base contributed from Mom.
In English: When allele1 doesn’t equal the allele that you got from Mom, put it in as the allele you got from Dad. This results in over 76,000 row changes. By the way, if I haven’t mentioned it, in the Update Query, the row that says Update To is the one where the update to your data is happening. So in the above example if Sharon’s allele1 doesn’t equal the one she got from Mom, that allele is then known to be the one from dad and is inserted in the correct place in a new table.
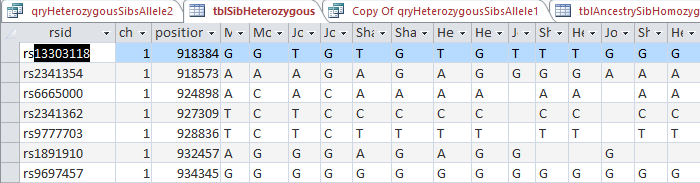
I check my updated table for good old rs13303118 and find:
So I think that it looks pretty good. The first line is now filled in with our Dad’s contributing base. Also all the applicable following lines out of 700,000. There are some situations there should be blanks. In the third line, my mom is AC and I am AC. That is the situation where it is not possible to know what base came from what parent. So the base for each of my contributing parent is left blank – meaning that it is unknown.
One Last Step: Looking for Patterns
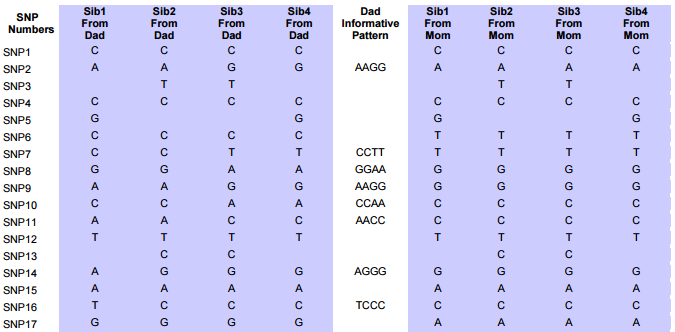
This is about as far as I’ve gotten and understood. The next issue that Whit Athey looked at in his paper were patterns. In his example there were 4 siblings tested, so more patterns. He added a column between the allele inherited from Dad and one from Mom called Dad informative pattern.
The idea is that there will be a pattern that lasts for a long time as we go down the results sequentially. These are the patterns of the segments that we inherit from our grandparents. Where the patterns change are the crossovers. Whit says to use those patterns to fill some of the missing letters. I haven’t started filling in the missing bases yet for a few reasons. One is that I’m not sure why I need to. In scanning the Athey paper there is a repetitive procedure of going back and forth between the data using the base from Dad’s side fill-in’s to help with the base from Mom’s side and then back again. First I’m not sure how to automate this yet. And if I could, how much better would the data be? I have quite a bit of data already. Once I get some answers of why I need to do this, I will continue on.
Here is a paragraph from the Athey Paper concerning the above Table 2:
Note the pattern of inheritance from Dad shown in Table 2 for the four siblings in the leftmost four columns. The first few rows show an AABB base pattern, but this gives way in about lines 12-13 to a new pattern, ABBB. Even though we only can see the pattern showing in some of the rows, these patterns persist over hundreds or thousands of SNPs, and can be assumed to exist also in the intervening rows where no pattern was discernable (and in the underlying sequence). Note that often there will be the same base in every location, a case of “accidental matching” which does not contribute to or detract from the pattern we are looking for. When two or more bases are different in a row, however, this represents an informative pattern—if any two are different, then since there are only two possible chromosomes contributing, it means we can see the chromosomal origins of the bases.
One of the reasons that I quote the above is to address the accidental matching where there was the same contributing parent base for each sibling. However, what I didn’t see addressed is that there are cases where that is not just accidental which I will discuss later.
Finding the crossovers
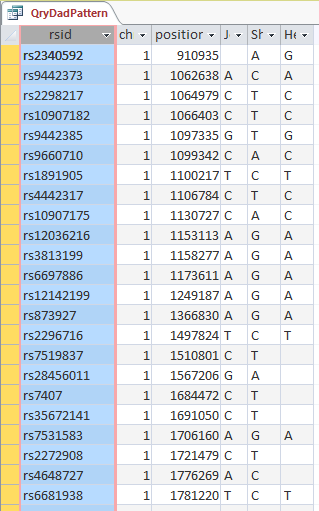
I do know the importance of finding the crossovers. I wrote a query in Access to cull out the patterns that Whit mentions.
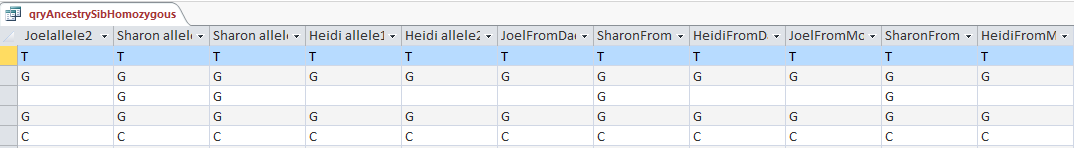
Above is my query in design view using the table that has Principles, 1, 2, and 3 already applied. This query basically filters out the situations where the 3 siblings have the same base. The thought is that if one sibling has one base that is different from one of the others, then the three siblings’ will not share the same base.
Above is the start of the results of the query. Note the XYX pattern. This should make it possible to fill in Heidi’s missing bases from Dad. It looks like multiple choice test answers, but I would add C, G, C, C, C and A in the last column for the bases that Heidi got from Dad. My homework assignment is to find a formula to fill in those letters so I don’t have to do it manually 10’s of thousands of times.
Another thing I want Access to do is find where the crossovers are. Here I scrolled down all the bases the my sisters and I had from Dad. I can see where the XYX pattern changes to XYY:
But there was a problem. the XYX pattern stopped at position 18,759,377 and the XYY pattern started at 23,288,828. That means we have a large area with no pattern. Exactly. That is the area of XXX pattern that I just queried out. That has to be the area where all three siblings match the same paternal grandparent.
Checking my results with m macneill’s work
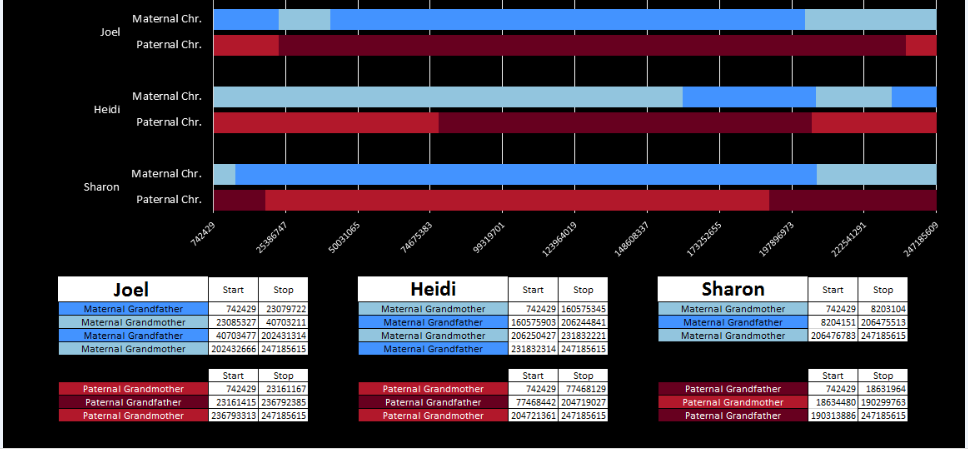
Fortunately, I have secret weapon. M MacNeill – prairielad_genealogy@hotmail.com has also been looking at my raw DNA using his own Excel spreadsheet method. Here is what he has for Chromosome 1:
Now just look at the first 3 red bars above. They represent my paternal side. The first break would be on Sharon’s bar – the third red bar from the top. The end of her dark red bar is at 18,631,964:
Look at Sharon’s bar in that region and then scan up the 3 red bars. There is an area where all three siblings match on the paternal grandmother side (lighter red).
That is my paternal XXX Pattern.
To satisfy my curiosity, I went back to my unfiltered/unqueried table at the spot that the first pattern changed from XYX to XXX. The end of the first base pattern from Dad is highlighted in blue.
Line 2 is a no-call. Line 3 is one of the random XXX matches in the XYX pattern area that Athey mentioned above. Note that I could not likely fill in line 4 with what I know as I don’t know if that should be AAA, AGA, or something else. Actually, I could fill in Heidi’s with an A. If her results are AAA or AGA, Heidi still gets the A from Dad. It is only Sharon’s base from Dad that I don’t know.
However, starting at CCC, it seems like it would make sense to fill in all the letters in the XXX pattern area – even if there is only one known base out of three.
Converting Build 37 to Build 36 positions
At the top of the Blog I had mentioned that AncestryDNA results were in Build 37. M MacNeill’s work is in Build 36. I really didn’t want to have to convert results and thought that I was being clever by using all AncestryDNA results. However, to compare to M MacNeill’s Map above or to Gedmatch results, I still have to convert positions. Hey, life is tough.
NCBI genome Remapping service
Fortunately there is a way to convert positions here.
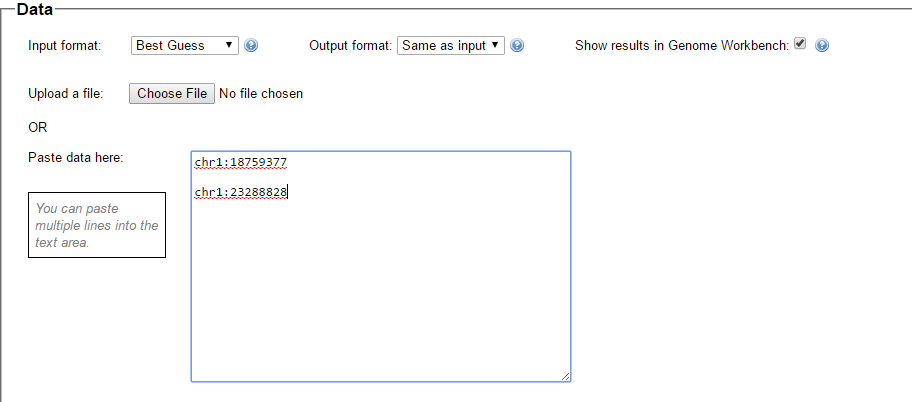
Assuming we are all homo sapiens, we select that choice and we select that we want to go from Build 37 to 36:
Here is the place to enter the data we want converted. It has to be in the format below – “chr1:” followed by the position number. There is also a place to upload a file which I haven’t tried.
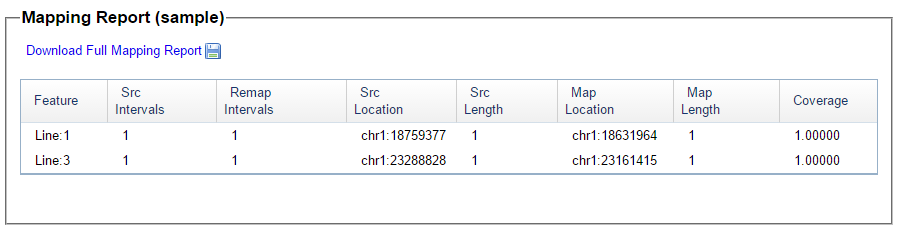
These are the 2 positions from my query where one pattern stopped and another started. Here is what they look like in Build 36 under Map Location:
These Build 36 position numbers match up perfectly with M MacNeill’s map positions which gives me some confidence. This is where I’ll end Part 1.
I have found 2 paternal crossover points. However, I have not yet figured out which siblings they belong to – unless I cheat and look at the MacNeill Map above. I can easily do the same thing and find the pattern changes for the maternal side. I have shown 2 crossovers, but all the others exist in my query for 23 chromosomes. I just haven’t looked for them yet.
Summary
- The Whit Athey Paper has been very helpful in phasing my raw DNA based on my mother and 2 siblings test results.
- M MacNeill has piqued an interest in raw DNA data that I never thought I would have
- M MacNeill’s Chromosome Maps are very helpful in checking my work
- MS Access appears to be a great tool to use to quickly phase a lot of raw DNA
- There is probably no way around DNA remapping or conversions
- I still need:
- An easy way to find all the crossover points
- A formula to fill in the various patterns
- A good reason to fill in those missing bases
- I have a lot more to learn about DNA phasing using raw DNA data